Scaling to large datasets¶
pandas provides data structures for in-memory analytics, which makes using pandas to analyze datasets that are larger than memory datasets somewhat tricky. Even datasets that are a sizable fraction of memory become unwieldy, as some pandas operations need to make intermediate copies.
This document provides a few recommendations for scaling your analysis to larger datasets. It’s a complement to Enhancing performance, which focuses on speeding up analysis for datasets that fit in memory.
But first, it’s worth considering not using pandas. pandas isn’t the right tool for all situations. If you’re working with very large datasets and a tool like PostgreSQL fits your needs, then you should probably be using that. Assuming you want or need the expressiveness and power of pandas, let’s carry on.
In [1]: import pandas as pd
In [2]: import numpy as np
Load less data¶
Suppose our raw dataset on disk has many columns:
id_0 name_0 x_0 y_0 id_1 name_1 x_1 ... name_8 x_8 y_8 id_9 name_9 x_9 y_9
timestamp ...
2000-01-01 00:00:00 1015 Michael -0.399453 0.095427 994 Frank -0.176842 ... Dan -0.315310 0.713892 1025 Victor -0.135779 0.346801
2000-01-01 00:01:00 969 Patricia 0.650773 -0.874275 1003 Laura 0.459153 ... Ursula 0.913244 -0.630308 1047 Wendy -0.886285 0.035852
2000-01-01 00:02:00 1016 Victor -0.721465 -0.584710 1046 Michael 0.524994 ... Ray -0.656593 0.692568 1064 Yvonne 0.070426 0.432047
2000-01-01 00:03:00 939 Alice -0.746004 -0.908008 996 Ingrid -0.414523 ... Jerry -0.958994 0.608210 978 Wendy 0.855949 -0.648988
2000-01-01 00:04:00 1017 Dan 0.919451 -0.803504 1048 Jerry -0.569235 ... Frank -0.577022 -0.409088 994 Bob -0.270132 0.335176
... ... ... ... ... ... ... ... ... ... ... ... ... ... ... ...
2000-12-30 23:56:00 999 Tim 0.162578 0.512817 973 Kevin -0.403352 ... Tim -0.380415 0.008097 1041 Charlie 0.191477 -0.599519
2000-12-30 23:57:00 970 Laura -0.433586 -0.600289 958 Oliver -0.966577 ... Zelda 0.971274 0.402032 1038 Ursula 0.574016 -0.930992
2000-12-30 23:58:00 1065 Edith 0.232211 -0.454540 971 Tim 0.158484 ... Alice -0.222079 -0.919274 1022 Dan 0.031345 -0.657755
2000-12-30 23:59:00 1019 Ingrid 0.322208 -0.615974 981 Hannah 0.607517 ... Sarah -0.424440 -0.117274 990 George -0.375530 0.563312
2000-12-31 00:00:00 937 Ursula -0.906523 0.943178 1018 Alice -0.564513 ... Jerry 0.236837 0.807650 985 Oliver 0.777642 0.783392
[525601 rows x 40 columns]
To load the columns we want, we have two options. Option 1 loads in all the data and then filters to what we need.
In [3]: columns = ["id_0", "name_0", "x_0", "y_0"]
In [4]: pd.read_parquet("timeseries_wide.parquet")[columns]
Out[4]:
id_0 name_0 x_0 y_0
timestamp
2000-01-01 00:00:00 1015 Michael -0.399453 0.095427
2000-01-01 00:01:00 969 Patricia 0.650773 -0.874275
2000-01-01 00:02:00 1016 Victor -0.721465 -0.584710
2000-01-01 00:03:00 939 Alice -0.746004 -0.908008
2000-01-01 00:04:00 1017 Dan 0.919451 -0.803504
... ... ... ... ...
2000-12-30 23:56:00 999 Tim 0.162578 0.512817
2000-12-30 23:57:00 970 Laura -0.433586 -0.600289
2000-12-30 23:58:00 1065 Edith 0.232211 -0.454540
2000-12-30 23:59:00 1019 Ingrid 0.322208 -0.615974
2000-12-31 00:00:00 937 Ursula -0.906523 0.943178
[525601 rows x 4 columns]
Option 2 only loads the columns we request.
In [5]: pd.read_parquet("timeseries_wide.parquet", columns=columns)
Out[5]:
id_0 name_0 x_0 y_0
timestamp
2000-01-01 00:00:00 1015 Michael -0.399453 0.095427
2000-01-01 00:01:00 969 Patricia 0.650773 -0.874275
2000-01-01 00:02:00 1016 Victor -0.721465 -0.584710
2000-01-01 00:03:00 939 Alice -0.746004 -0.908008
2000-01-01 00:04:00 1017 Dan 0.919451 -0.803504
... ... ... ... ...
2000-12-30 23:56:00 999 Tim 0.162578 0.512817
2000-12-30 23:57:00 970 Laura -0.433586 -0.600289
2000-12-30 23:58:00 1065 Edith 0.232211 -0.454540
2000-12-30 23:59:00 1019 Ingrid 0.322208 -0.615974
2000-12-31 00:00:00 937 Ursula -0.906523 0.943178
[525601 rows x 4 columns]
If we were to measure the memory usage of the two calls, we’d see that specifying
columns uses about 1/10th the memory in this case.
With pandas.read_csv(), you can specify usecols to limit the columns
read into memory. Not all file formats that can be read by pandas provide an option
to read a subset of columns.
Use efficient datatypes¶
The default pandas data types are not the most memory efficient. This is especially true for text data columns with relatively few unique values (commonly referred to as “low-cardinality” data). By using more efficient data types, you can store larger datasets in memory.
In [6]: ts = pd.read_parquet("timeseries.parquet")
In [7]: ts
Out[7]:
id name x y
timestamp
2000-01-01 00:00:00 1029 Michael 0.278837 0.247932
2000-01-01 00:00:30 1010 Patricia 0.077144 0.490260
2000-01-01 00:01:00 1001 Victor 0.214525 0.258635
2000-01-01 00:01:30 1018 Alice -0.646866 0.822104
2000-01-01 00:02:00 991 Dan 0.902389 0.466665
... ... ... ... ...
2000-12-30 23:58:00 992 Sarah 0.721155 0.944118
2000-12-30 23:58:30 1007 Ursula 0.409277 0.133227
2000-12-30 23:59:00 1009 Hannah -0.452802 0.184318
2000-12-30 23:59:30 978 Kevin -0.904728 -0.179146
2000-12-31 00:00:00 973 Ingrid -0.370763 -0.794667
[1051201 rows x 4 columns]
Now, let’s inspect the data types and memory usage to see where we should focus our attention.
In [8]: ts.dtypes
Out[8]:
id int64
name object
x float64
y float64
dtype: object
In [9]: ts.memory_usage(deep=True) # memory usage in bytes
Out[9]:
Index 8409608
id 8409608
name 65537768
x 8409608
y 8409608
dtype: int64
The name column is taking up much more memory than any other. It has just a
few unique values, so it’s a good candidate for converting to a
Categorical. With a Categorical, we store each unique name once and use
space-efficient integers to know which specific name is used in each row.
In [10]: ts2 = ts.copy()
In [11]: ts2["name"] = ts2["name"].astype("category")
In [12]: ts2.memory_usage(deep=True)
Out[12]:
Index 8409608
id 8409608
name 1053894
x 8409608
y 8409608
dtype: int64
We can go a bit further and downcast the numeric columns to their smallest types
using pandas.to_numeric().
In [13]: ts2["id"] = pd.to_numeric(ts2["id"], downcast="unsigned")
In [14]: ts2[["x", "y"]] = ts2[["x", "y"]].apply(pd.to_numeric, downcast="float")
In [15]: ts2.dtypes
Out[15]:
id uint16
name category
x float32
y float32
dtype: object
In [16]: ts2.memory_usage(deep=True)
Out[16]:
Index 8409608
id 2102402
name 1053894
x 4204804
y 4204804
dtype: int64
In [17]: reduction = ts2.memory_usage(deep=True).sum() / ts.memory_usage(deep=True).sum()
In [18]: print(f"{reduction:0.2f}")
0.20
In all, we’ve reduced the in-memory footprint of this dataset to 1/5 of its original size.
See Categorical data for more on Categorical and dtypes
for an overview of all of pandas’ dtypes.
Use chunking¶
Some workloads can be achieved with chunking: splitting a large problem like “convert this directory of CSVs to parquet” into a bunch of small problems (“convert this individual CSV file into a Parquet file. Now repeat that for each file in this directory.”). As long as each chunk fits in memory, you can work with datasets that are much larger than memory.
Note
Chunking works well when the operation you’re performing requires zero or minimal coordination between chunks. For more complicated workflows, you’re better off using another library.
Suppose we have an even larger “logical dataset” on disk that’s a directory of parquet files. Each file in the directory represents a different year of the entire dataset.
data
└── timeseries
├── ts-00.parquet
├── ts-01.parquet
├── ts-02.parquet
├── ts-03.parquet
├── ts-04.parquet
├── ts-05.parquet
├── ts-06.parquet
├── ts-07.parquet
├── ts-08.parquet
├── ts-09.parquet
├── ts-10.parquet
└── ts-11.parquet
Now we’ll implement an out-of-core value_counts. The peak memory usage of this
workflow is the single largest chunk, plus a small series storing the unique value
counts up to this point. As long as each individual file fits in memory, this will
work for arbitrary-sized datasets.
In [19]: %%time
....: files = pathlib.Path("data/timeseries/").glob("ts*.parquet")
....: counts = pd.Series(dtype=int)
....: for path in files:
....: df = pd.read_parquet(path)
....: counts = counts.add(df["name"].value_counts(), fill_value=0)
....: counts.astype(int)
....:
CPU times: user 1.36 s, sys: 83.2 ms, total: 1.45 s
Wall time: 1.13 s
Out[19]:
Alice 229802
Bob 229211
Charlie 229303
Dan 230621
Edith 230349
...
Victor 230502
Wendy 230038
Xavier 229553
Yvonne 228766
Zelda 229909
Length: 26, dtype: int64
Some readers, like pandas.read_csv(), offer parameters to control the
chunksize when reading a single file.
Manually chunking is an OK option for workflows that don’t
require too sophisticated of operations. Some operations, like groupby, are
much harder to do chunkwise. In these cases, you may be better switching to a
different library that implements these out-of-core algorithms for you.
Use other libraries¶
pandas is just one library offering a DataFrame API. Because of its popularity, pandas’ API has become something of a standard that other libraries implement. The pandas documentation maintains a list of libraries implementing a DataFrame API in our ecosystem page.
For example, Dask, a parallel computing library, has dask.dataframe, a pandas-like API for working with larger than memory datasets in parallel. Dask can use multiple threads or processes on a single machine, or a cluster of machines to process data in parallel.
We’ll import dask.dataframe and notice that the API feels similar to pandas.
We can use Dask’s read_parquet function, but provide a globstring of files to read in.
In [20]: import dask.dataframe as dd
In [21]: ddf = dd.read_parquet("data/timeseries/ts*.parquet", engine="pyarrow")
In [22]: ddf
Out[22]:
Dask DataFrame Structure:
id name x y
npartitions=12
int64 object float64 float64
... ... ... ...
... ... ... ... ...
... ... ... ...
... ... ... ...
Dask Name: read-parquet, 1 graph layer
Inspecting the ddf object, we see a few things
There are familiar attributes like
.columnsand.dtypesThere are familiar methods like
.groupby,.sum, etc.There are new attributes like
.npartitionsand.divisions
The partitions and divisions are how Dask parallelizes computation. A Dask DataFrame is made up of many pandas DataFrames. A single method call on a Dask DataFrame ends up making many pandas method calls, and Dask knows how to coordinate everything to get the result.
In [23]: ddf.columns
Out[23]: Index(['id', 'name', 'x', 'y'], dtype='object')
In [24]: ddf.dtypes
Out[24]:
id int64
name object
x float64
y float64
dtype: object
In [25]: ddf.npartitions
Out[25]: 12
One major difference: the dask.dataframe API is lazy. If you look at the
repr above, you’ll notice that the values aren’t actually printed out; just the
column names and dtypes. That’s because Dask hasn’t actually read the data yet.
Rather than executing immediately, doing operations build up a task graph.
In [26]: ddf
Out[26]:
Dask DataFrame Structure:
id name x y
npartitions=12
int64 object float64 float64
... ... ... ...
... ... ... ... ...
... ... ... ...
... ... ... ...
Dask Name: read-parquet, 1 graph layer
In [27]: ddf["name"]
Out[27]:
Dask Series Structure:
npartitions=12
object
...
...
...
...
Name: name, dtype: object
Dask Name: getitem, 2 graph layers
In [28]: ddf["name"].value_counts()
Out[28]:
Dask Series Structure:
npartitions=1
int64
...
Name: name, dtype: int64
Dask Name: value-counts-agg, 4 graph layers
Each of these calls is instant because the result isn’t being computed yet.
We’re just building up a list of computation to do when someone needs the
result. Dask knows that the return type of a pandas.Series.value_counts
is a pandas Series with a certain dtype and a certain name. So the Dask version
returns a Dask Series with the same dtype and the same name.
To get the actual result you can call .compute().
In [29]: %time ddf["name"].value_counts().compute()
CPU times: user 1.54 s, sys: 211 ms, total: 1.75 s
Wall time: 1.2 s
Out[29]:
Laura 230906
Ingrid 230838
Kevin 230698
Dan 230621
Frank 230595
...
Ray 229603
Xavier 229553
Charlie 229303
Bob 229211
Yvonne 228766
Name: name, Length: 26, dtype: int64
At that point, you get back the same thing you’d get with pandas, in this case
a concrete pandas Series with the count of each name.
Calling .compute causes the full task graph to be executed. This includes
reading the data, selecting the columns, and doing the value_counts. The
execution is done in parallel where possible, and Dask tries to keep the
overall memory footprint small. You can work with datasets that are much larger
than memory, as long as each partition (a regular pandas DataFrame) fits in memory.
By default, dask.dataframe operations use a threadpool to do operations in
parallel. We can also connect to a cluster to distribute the work on many
machines. In this case we’ll connect to a local “cluster” made up of several
processes on this single machine.
>>> from dask.distributed import Client, LocalCluster
>>> cluster = LocalCluster()
>>> client = Client(cluster)
>>> client
<Client: 'tcp://127.0.0.1:53349' processes=4 threads=8, memory=17.18 GB>
Once this client is created, all of Dask’s computation will take place on
the cluster (which is just processes in this case).
Dask implements the most used parts of the pandas API. For example, we can do a familiar groupby aggregation.
In [30]: %time ddf.groupby("name")[["x", "y"]].mean().compute().head()
CPU times: user 9.52 s, sys: 1.79 s, total: 11.3 s
Wall time: 8.45 s
Out[30]:
x y
name
Alice 0.000086 -0.001170
Bob -0.000843 -0.000799
Charlie 0.000564 -0.000038
Dan 0.000584 0.000818
Edith -0.000116 -0.000044
The grouping and aggregation is done out-of-core and in parallel.
When Dask knows the divisions of a dataset, certain optimizations are
possible. When reading parquet datasets written by dask, the divisions will be
known automatically. In this case, since we created the parquet files manually,
we need to supply the divisions manually.
In [31]: N = 12
In [32]: starts = [f"20{i:>02d}-01-01" for i in range(N)]
In [33]: ends = [f"20{i:>02d}-12-13" for i in range(N)]
In [34]: divisions = tuple(pd.to_datetime(starts)) + (pd.Timestamp(ends[-1]),)
In [35]: ddf.divisions = divisions
In [36]: ddf
Out[36]:
Dask DataFrame Structure:
id name x y
npartitions=12
2000-01-01 int64 object float64 float64
2001-01-01 ... ... ... ...
... ... ... ... ...
2011-01-01 ... ... ... ...
2011-12-13 ... ... ... ...
Dask Name: read-parquet, 1 graph layer
Now we can do things like fast random access with .loc.
In [37]: ddf.loc["2002-01-01 12:01":"2002-01-01 12:05"].compute()
Out[37]:
id name x y
timestamp
2002-01-01 12:01:00 983 Laura 0.243985 -0.079392
2002-01-01 12:02:00 1001 Laura -0.523119 -0.226026
2002-01-01 12:03:00 1059 Oliver 0.612886 0.405680
2002-01-01 12:04:00 993 Kevin 0.451977 0.332947
2002-01-01 12:05:00 1014 Yvonne -0.948681 0.361748
Dask knows to just look in the 3rd partition for selecting values in 2002. It doesn’t need to look at any other data.
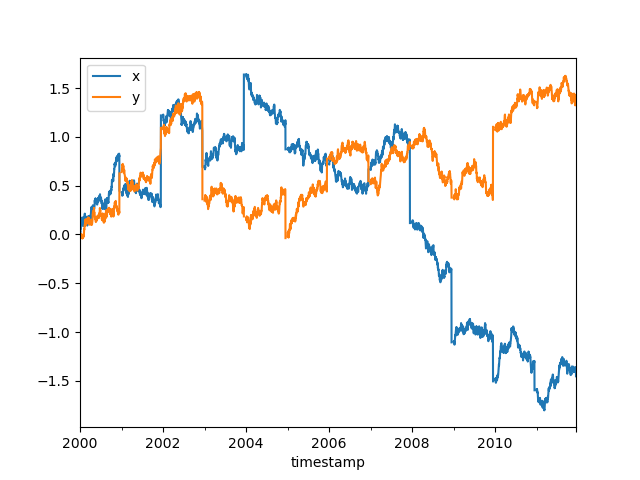
Many workflows involve a large amount of data and processing it in a way that
reduces the size to something that fits in memory. In this case, we’ll resample
to daily frequency and take the mean. Once we’ve taken the mean, we know the
results will fit in memory, so we can safely call compute without running
out of memory. At that point it’s just a regular pandas object.
In [38]: ddf[["x", "y"]].resample("1D").mean().cumsum().compute().plot()
Out[38]: <AxesSubplot:xlabel='timestamp'>
These Dask examples have all be done using multiple processes on a single machine. Dask can be deployed on a cluster to scale up to even larger datasets.
You see more dask examples at https://examples.dask.org.