Version 0.13.0 (January 3, 2014)#
This is a major release from 0.12.0 and includes a number of API changes, several new features and enhancements along with a large number of bug fixes.
Highlights include:
support for a new index type
Float64Index, and other Indexing enhancementsHDFStorehas a new string based syntax for query specificationsupport for new methods of interpolation
updated
timedeltaoperationsa new string manipulation method
extractNanosecond support for Offsets
isinfor DataFrames
Several experimental features are added, including:
new
eval/querymethods for expression evaluationsupport for
msgpackserializationan i/o interface to Google’s
BigQuery
Their are several new or updated docs sections including:
Comparison with SQL, which should be useful for those familiar with SQL but still learning pandas.
Comparison with R, idiom translations from R to pandas.
Enhancing Performance, ways to enhance pandas performance with
eval/query.
Warning
In 0.13.0 Series has internally been refactored to no longer sub-class ndarray
but instead subclass NDFrame, similar to the rest of the pandas containers. This should be
a transparent change with only very limited API implications. See Internal Refactoring
API changes#
read_excelnow supports an integer in itssheetnameargument giving the index of the sheet to read in (GH 4301).Text parser now treats anything that reads like inf (“inf”, “Inf”, “-Inf”, “iNf”, etc.) as infinity. (GH 4220, GH 4219), affecting
read_table,read_csv, etc.pandasnow is Python 2/3 compatible without the need for 2to3 thanks to @jtratner. As a result, pandas now uses iterators more extensively. This also led to the introduction of substantive parts of the Benjamin Peterson’ssixlibrary into compat. (GH 4384, GH 4375, GH 4372)pandas.util.compatandpandas.util.py3compathave been merged intopandas.compat.pandas.compatnow includes many functions allowing 2/3 compatibility. It contains both list and iterator versions of range, filter, map and zip, plus other necessary elements for Python 3 compatibility.lmap,lzip,lrangeandlfilterall produce lists instead of iterators, for compatibility withnumpy, subscripting andpandasconstructors.(GH 4384, GH 4375, GH 4372)Series.getwith negative indexers now returns the same as[](GH 4390)Changes to how
IndexandMultiIndexhandle metadata (levels,labels, andnames) (GH 4039):# previously, you would have set levels or labels directly >>> pd.index.levels = [[1, 2, 3, 4], [1, 2, 4, 4]] # now, you use the set_levels or set_labels methods >>> index = pd.index.set_levels([[1, 2, 3, 4], [1, 2, 4, 4]]) # similarly, for names, you can rename the object # but setting names is not deprecated >>> index = pd.index.set_names(["bob", "cranberry"]) # and all methods take an inplace kwarg - but return None >>> pd.index.set_names(["bob", "cranberry"], inplace=True)
All division with
NDFrameobjects is now truedivision, regardless of the future import. This means that operating on pandas objects will by default use floating point division, and return a floating point dtype. You can use//andfloordivto do integer division.Integer division
In [3]: arr = np.array([1, 2, 3, 4]) In [4]: arr2 = np.array([5, 3, 2, 1]) In [5]: arr / arr2 Out[5]: array([0, 0, 1, 4]) In [6]: pd.Series(arr) // pd.Series(arr2) Out[6]: 0 0 1 0 2 1 3 4 dtype: int64
True Division
In [7]: pd.Series(arr) / pd.Series(arr2) # no future import required Out[7]: 0 0.200000 1 0.666667 2 1.500000 3 4.000000 dtype: float64
Infer and downcast dtype if
downcast='infer'is passed tofillna/ffill/bfill(GH 4604)__nonzero__for all NDFrame objects, will now raise aValueError, this reverts back to (GH 1073, GH 4633) behavior. See gotchas for a more detailed discussion.This prevents doing boolean comparison on entire pandas objects, which is inherently ambiguous. These all will raise a
ValueError.>>> df = pd.DataFrame({'A': np.random.randn(10), ... 'B': np.random.randn(10), ... 'C': pd.date_range('20130101', periods=10) ... }) ... >>> if df: ... pass ... Traceback (most recent call last): ... ValueError: The truth value of a DataFrame is ambiguous. Use a.empty, a.bool(), a.item(), a.any() or a.all(). >>> df1 = df >>> df2 = df >>> df1 and df2 Traceback (most recent call last): ... ValueError: The truth value of a DataFrame is ambiguous. Use a.empty, a.bool(), a.item(), a.any() or a.all(). >>> d = [1, 2, 3] >>> s1 = pd.Series(d) >>> s2 = pd.Series(d) >>> s1 and s2 Traceback (most recent call last): ... ValueError: The truth value of a DataFrame is ambiguous. Use a.empty, a.bool(), a.item(), a.any() or a.all().
Added the
.bool()method toNDFrameobjects to facilitate evaluating of single-element boolean Series:>>> pd.Series([True]).bool() True >>> pd.Series([False]).bool() False >>> pd.DataFrame([[True]]).bool() True >>> pd.DataFrame([[False]]).bool() False
All non-Index NDFrames (
Series,DataFrame,Panel,Panel4D,SparsePanel, etc.), now support the entire set of arithmetic operators and arithmetic flex methods (add, sub, mul, etc.).SparsePaneldoes not supportpowormodwith non-scalars. (GH 3765)SeriesandDataFramenow have amode()method to calculate the statistical mode(s) by axis/Series. (GH 5367)Chained assignment will now by default warn if the user is assigning to a copy. This can be changed with the option
mode.chained_assignment, allowed options areraise/warn/None. See the docs.In [1]: dfc = pd.DataFrame({'A': ['aaa', 'bbb', 'ccc'], 'B': [1, 2, 3]}) In [2]: pd.set_option('chained_assignment', 'warn')
The following warning / exception will show if this is attempted.
In [3]: dfc.loc[0]['A'] = 1111
Traceback (most recent call last) ... SettingWithCopyWarning: A value is trying to be set on a copy of a slice from a DataFrame. Try using .loc[row_index,col_indexer] = value instead
Here is the correct method of assignment.
In [4]: dfc.loc[0, 'A'] = 11 In [5]: dfc Out[5]: A B 0 11 1 1 bbb 2 2 ccc 3
Panel.reindexhas the following call signaturePanel.reindex(items=None, major_axis=None, minor_axis=None, **kwargs)to conform with other
NDFrameobjects. See Internal Refactoring for more information.
Series.argminandSeries.argmaxare now aliased toSeries.idxminandSeries.idxmax. These return the index of themin or max element respectively. Prior to 0.13.0 these would return the position of the min / max element. (GH 6214)
Prior version deprecations/changes#
These were announced changes in 0.12 or prior that are taking effect as of 0.13.0
Remove deprecated
Factor(GH 3650)Remove deprecated
set_printoptions/reset_printoptions(GH 3046)Remove deprecated
_verbose_info(GH 3215)Remove deprecated
read_clipboard/to_clipboard/ExcelFile/ExcelWriterfrompandas.io.parsers(GH 3717) These are available as functions in the main pandas namespace (e.g.pd.read_clipboard)default for
tupleize_colsis nowFalsefor bothto_csvandread_csv. Fair warning in 0.12 (GH 3604)default for
display.max_seq_lenis now 100 rather thanNone. This activates truncated display (”…”) of long sequences in various places. (GH 3391)
Deprecations#
Deprecated in 0.13.0
deprecated
iterkv, which will be removed in a future release (this was an alias of iteritems used to bypass2to3’s changes). (GH 4384, GH 4375, GH 4372)deprecated the string method
match, whose role is now performed more idiomatically byextract. In a future release, the default behavior ofmatchwill change to become analogous tocontains, which returns a boolean indexer. (Their distinction is strictness:matchrelies onre.matchwhilecontainsrelies onre.search.) In this release, the deprecated behavior is the default, but the new behavior is available through the keyword argumentas_indexer=True.
Indexing API changes#
Prior to 0.13, it was impossible to use a label indexer (.loc/.ix) to set a value that
was not contained in the index of a particular axis. (GH 2578). See the docs
In the Series case this is effectively an appending operation
In [6]: s = pd.Series([1, 2, 3])
In [7]: s
Out[7]:
0 1
1 2
2 3
dtype: int64
In [8]: s[5] = 5.
In [9]: s
Out[9]:
0 1.0
1 2.0
2 3.0
5 5.0
dtype: float64
In [10]: dfi = pd.DataFrame(np.arange(6).reshape(3, 2),
....: columns=['A', 'B'])
....:
In [11]: dfi
Out[11]:
A B
0 0 1
1 2 3
2 4 5
This would previously KeyError
In [12]: dfi.loc[:, 'C'] = dfi.loc[:, 'A']
In [13]: dfi
Out[13]:
A B C
0 0 1 0
1 2 3 2
2 4 5 4
This is like an append operation.
In [14]: dfi.loc[3] = 5
In [15]: dfi
Out[15]:
A B C
0 0 1 0
1 2 3 2
2 4 5 4
3 5 5 5
A Panel setting operation on an arbitrary axis aligns the input to the Panel
In [20]: p = pd.Panel(np.arange(16).reshape(2, 4, 2),
....: items=['Item1', 'Item2'],
....: major_axis=pd.date_range('2001/1/12', periods=4),
....: minor_axis=['A', 'B'], dtype='float64')
....:
In [21]: p
Out[21]:
<class 'pandas.core.panel.Panel'>
Dimensions: 2 (items) x 4 (major_axis) x 2 (minor_axis)
Items axis: Item1 to Item2
Major_axis axis: 2001-01-12 00:00:00 to 2001-01-15 00:00:00
Minor_axis axis: A to B
In [22]: p.loc[:, :, 'C'] = pd.Series([30, 32], index=p.items)
In [23]: p
Out[23]:
<class 'pandas.core.panel.Panel'>
Dimensions: 2 (items) x 4 (major_axis) x 3 (minor_axis)
Items axis: Item1 to Item2
Major_axis axis: 2001-01-12 00:00:00 to 2001-01-15 00:00:00
Minor_axis axis: A to C
In [24]: p.loc[:, :, 'C']
Out[24]:
Item1 Item2
2001-01-12 30.0 32.0
2001-01-13 30.0 32.0
2001-01-14 30.0 32.0
2001-01-15 30.0 32.0
Float64Index API change#
Added a new index type,
Float64Index. This will be automatically created when passing floating values in index creation. This enables a pure label-based slicing paradigm that makes[],ix,locfor scalar indexing and slicing work exactly the same. (GH 263)Construction is by default for floating type values.
In [16]: index = pd.Index([1.5, 2, 3, 4.5, 5]) In [17]: index Out[17]: Index([1.5, 2.0, 3.0, 4.5, 5.0], dtype='float64') In [18]: s = pd.Series(range(5), index=index) In [19]: s Out[19]: 1.5 0 2.0 1 3.0 2 4.5 3 5.0 4 dtype: int64
Scalar selection for
[],.ix,.locwill always be label based. An integer will match an equal float index (e.g.3is equivalent to3.0)In [20]: s[3] Out[20]: 2 In [21]: s.loc[3] Out[21]: 2
The only positional indexing is via
ilocIn [22]: s.iloc[3] Out[22]: 3
A scalar index that is not found will raise
KeyErrorSlicing is ALWAYS on the values of the index, for
[],ix,locand ALWAYS positional withilocIn [23]: s[2:4] Out[23]: 2.0 1 3.0 2 dtype: int64 In [24]: s.loc[2:4] Out[24]: 2.0 1 3.0 2 dtype: int64 In [25]: s.iloc[2:4] Out[25]: 3.0 2 4.5 3 dtype: int64
In float indexes, slicing using floats are allowed
In [26]: s[2.1:4.6] Out[26]: 3.0 2 4.5 3 dtype: int64 In [27]: s.loc[2.1:4.6] Out[27]: 3.0 2 4.5 3 dtype: int64
Indexing on other index types are preserved (and positional fallback for
[],ix), with the exception, that floating point slicing on indexes on nonFloat64Indexwill now raise aTypeError.In [1]: pd.Series(range(5))[3.5] TypeError: the label [3.5] is not a proper indexer for this index type (Int64Index) In [1]: pd.Series(range(5))[3.5:4.5] TypeError: the slice start [3.5] is not a proper indexer for this index type (Int64Index)
Using a scalar float indexer will be deprecated in a future version, but is allowed for now.
In [3]: pd.Series(range(5))[3.0] Out[3]: 3
HDFStore API changes#
Query Format Changes. A much more string-like query format is now supported. See the docs.
In [28]: path = 'test.h5' In [29]: dfq = pd.DataFrame(np.random.randn(10, 4), ....: columns=list('ABCD'), ....: index=pd.date_range('20130101', periods=10)) ....: In [30]: dfq.to_hdf(path, 'dfq', format='table', data_columns=True)
Use boolean expressions, with in-line function evaluation.
In [31]: pd.read_hdf(path, 'dfq', ....: where="index>Timestamp('20130104') & columns=['A', 'B']") ....: Out[31]: A B 2013-01-05 -0.424972 0.567020 2013-01-06 -0.673690 0.113648 2013-01-07 0.404705 0.577046 2013-01-08 -0.370647 -1.157892 2013-01-09 1.075770 -0.109050 2013-01-10 0.357021 -0.674600
Use an inline column reference
In [32]: pd.read_hdf(path, 'dfq', ....: where="A>0 or C>0") ....: Out[32]: A B C D 2013-01-01 0.469112 -0.282863 -1.509059 -1.135632 2013-01-02 1.212112 -0.173215 0.119209 -1.044236 2013-01-04 0.721555 -0.706771 -1.039575 0.271860 2013-01-05 -0.424972 0.567020 0.276232 -1.087401 2013-01-07 0.404705 0.577046 -1.715002 -1.039268 2013-01-09 1.075770 -0.109050 1.643563 -1.469388 2013-01-10 0.357021 -0.674600 -1.776904 -0.968914
the
formatkeyword now replaces thetablekeyword; allowed values arefixed(f)ortable(t)the same defaults as prior < 0.13.0 remain, e.g.putimpliesfixedformat andappendimpliestableformat. This default format can be set as an option by settingio.hdf.default_format.In [33]: path = 'test.h5' In [34]: df = pd.DataFrame(np.random.randn(10, 2)) In [35]: df.to_hdf(path, 'df_table', format='table') In [36]: df.to_hdf(path, 'df_table2', append=True) In [37]: df.to_hdf(path, 'df_fixed') In [38]: with pd.HDFStore(path) as store: ....: print(store) ....: <class 'pandas.io.pytables.HDFStore'> File path: test.h5
Significant table writing performance improvements
handle a passed
Seriesin table format (GH 4330)can now serialize a
timedelta64[ns]dtype in a table (GH 3577), See the docs.added an
is_openproperty to indicate if the underlying file handle is_open; a closed store will now report ‘CLOSED’ when viewing the store (rather than raising an error) (GH 4409)a close of a
HDFStorenow will close that instance of theHDFStorebut will only close the actual file if the ref count (byPyTables) w.r.t. all of the open handles are 0. Essentially you have a local instance ofHDFStorereferenced by a variable. Once you close it, it will report closed. Other references (to the same file) will continue to operate until they themselves are closed. Performing an action on a closed file will raiseClosedFileErrorIn [39]: path = 'test.h5' In [40]: df = pd.DataFrame(np.random.randn(10, 2)) In [41]: store1 = pd.HDFStore(path) In [42]: store2 = pd.HDFStore(path) In [43]: store1.append('df', df) In [44]: store2.append('df2', df) In [45]: store1 Out[45]: <class 'pandas.io.pytables.HDFStore'> File path: test.h5 In [46]: store2 Out[46]: <class 'pandas.io.pytables.HDFStore'> File path: test.h5 In [47]: store1.close() In [48]: store2 Out[48]: <class 'pandas.io.pytables.HDFStore'> File path: test.h5 In [49]: store2.close() In [50]: store2 Out[50]: <class 'pandas.io.pytables.HDFStore'> File path: test.h5
removed the
_quietattribute, replace by aDuplicateWarningif retrieving duplicate rows from a table (GH 4367)removed the
warnargument fromopen. Instead aPossibleDataLossErrorexception will be raised if you try to usemode='w'with an OPEN file handle (GH 4367)allow a passed locations array or mask as a
wherecondition (GH 4467). See the docs for an example.add the keyword
dropna=Truetoappendto change whether ALL nan rows are not written to the store (default isTrue, ALL nan rows are NOT written), also settable via the optionio.hdf.dropna_table(GH 4625)pass through store creation arguments; can be used to support in-memory stores
DataFrame repr changes#
The HTML and plain text representations of DataFrame now show
a truncated view of the table once it exceeds a certain size, rather
than switching to the short info view (GH 4886, GH 5550).
This makes the representation more consistent as small DataFrames get
larger.
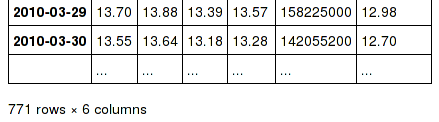
To get the info view, call DataFrame.info(). If you prefer the
info view as the repr for large DataFrames, you can set this by running
set_option('display.large_repr', 'info').
Enhancements#
df.to_clipboard()learned a newexcelkeyword that let’s you paste df data directly into excel (enabled by default). (GH 5070).read_htmlnow raises aURLErrorinstead of catching and raising aValueError(GH 4303, GH 4305)Added a test for
read_clipboard()andto_clipboard()(GH 4282)Clipboard functionality now works with PySide (GH 4282)
Added a more informative error message when plot arguments contain overlapping color and style arguments (GH 4402)
to_dictnow takesrecordsas a possible out type. Returns an array of column-keyed dictionaries. (GH 4936)NaNhanding in get_dummies (GH 4446) withdummy_na# previously, nan was erroneously counted as 2 here # now it is not counted at all In [51]: pd.get_dummies([1, 2, np.nan]) Out[51]: 1.0 2.0 0 True False 1 False True 2 False False # unless requested In [52]: pd.get_dummies([1, 2, np.nan], dummy_na=True) Out[52]: 1.0 2.0 NaN 0 True False False 1 False True False 2 False False True
timedelta64[ns]operations. See the docs.Warning
Most of these operations require
numpy >= 1.7Using the new top-level
to_timedelta, you can convert a scalar or array from the standard timedelta format (produced byto_csv) into a timedelta type (np.timedelta64innanoseconds).In [53]: pd.to_timedelta('1 days 06:05:01.00003') Out[53]: Timedelta('1 days 06:05:01.000030') In [54]: pd.to_timedelta('15.5us') Out[54]: Timedelta('0 days 00:00:00.000015500') In [55]: pd.to_timedelta(['1 days 06:05:01.00003', '15.5us', 'nan']) Out[55]: TimedeltaIndex(['1 days 06:05:01.000030', '0 days 00:00:00.000015500', NaT], dtype='timedelta64[ns]', freq=None) In [56]: pd.to_timedelta(np.arange(5), unit='s') Out[56]: TimedeltaIndex(['0 days 00:00:00', '0 days 00:00:01', '0 days 00:00:02', '0 days 00:00:03', '0 days 00:00:04'], dtype='timedelta64[ns]', freq=None) In [57]: pd.to_timedelta(np.arange(5), unit='d') Out[57]: TimedeltaIndex(['0 days', '1 days', '2 days', '3 days', '4 days'], dtype='timedelta64[ns]', freq=None)
A Series of dtype
timedelta64[ns]can now be divided by anothertimedelta64[ns]object, or astyped to yield afloat64dtyped Series. This is frequency conversion. See the docs for the docs.In [58]: import datetime In [59]: td = pd.Series(pd.date_range('20130101', periods=4)) - pd.Series( ....: pd.date_range('20121201', periods=4)) ....: In [60]: td[2] += np.timedelta64(datetime.timedelta(minutes=5, seconds=3)) In [61]: td[3] = np.nan In [62]: td Out[62]: 0 31 days 00:00:00 1 31 days 00:00:00 2 31 days 00:05:03 3 NaT dtype: timedelta64[ns] # to days In [63]: td / np.timedelta64(1, 'D') Out[63]: 0 31.000000 1 31.000000 2 31.003507 3 NaN dtype: float64 In [64]: td.astype('timedelta64[D]') --------------------------------------------------------------------------- ValueError Traceback (most recent call last) Cell In[64], line 1 ----> 1 td.astype('timedelta64[D]') File ~/work/pandas/pandas/pandas/core/generic.py:6532, in NDFrame.astype(self, dtype, copy, errors) 6528 results = [ser.astype(dtype, copy=copy) for _, ser in self.items()] 6530 else: 6531 # else, only a single dtype is given -> 6532 new_data = self._mgr.astype(dtype=dtype, copy=copy, errors=errors) 6533 res = self._constructor_from_mgr(new_data, axes=new_data.axes) 6534 return res.__finalize__(self, method="astype") File ~/work/pandas/pandas/pandas/core/internals/managers.py:414, in BaseBlockManager.astype(self, dtype, copy, errors) 411 elif using_copy_on_write(): 412 copy = False --> 414 return self.apply( 415 "astype", 416 dtype=dtype, 417 copy=copy, 418 errors=errors, 419 using_cow=using_copy_on_write(), 420 ) File ~/work/pandas/pandas/pandas/core/internals/managers.py:354, in BaseBlockManager.apply(self, f, align_keys, **kwargs) 352 applied = b.apply(f, **kwargs) 353 else: --> 354 applied = getattr(b, f)(**kwargs) 355 result_blocks = extend_blocks(applied, result_blocks) 357 out = type(self).from_blocks(result_blocks, self.axes) File ~/work/pandas/pandas/pandas/core/internals/blocks.py:616, in Block.astype(self, dtype, copy, errors, using_cow) 596 """ 597 Coerce to the new dtype. 598 (...) 612 Block 613 """ 614 values = self.values --> 616 new_values = astype_array_safe(values, dtype, copy=copy, errors=errors) 618 new_values = maybe_coerce_values(new_values) 620 refs = None File ~/work/pandas/pandas/pandas/core/dtypes/astype.py:238, in astype_array_safe(values, dtype, copy, errors) 235 dtype = dtype.numpy_dtype 237 try: --> 238 new_values = astype_array(values, dtype, copy=copy) 239 except (ValueError, TypeError): 240 # e.g. _astype_nansafe can fail on object-dtype of strings 241 # trying to convert to float 242 if errors == "ignore": File ~/work/pandas/pandas/pandas/core/dtypes/astype.py:180, in astype_array(values, dtype, copy) 176 return values 178 if not isinstance(values, np.ndarray): 179 # i.e. ExtensionArray --> 180 values = values.astype(dtype, copy=copy) 182 else: 183 values = _astype_nansafe(values, dtype, copy=copy) File ~/work/pandas/pandas/pandas/core/arrays/timedeltas.py:380, in TimedeltaArray.astype(self, dtype, copy) 376 return type(self)._simple_new( 377 res_values, dtype=res_values.dtype, freq=self.freq 378 ) 379 else: --> 380 raise ValueError( 381 f"Cannot convert from {self.dtype} to {dtype}. " 382 "Supported resolutions are 's', 'ms', 'us', 'ns'" 383 ) 385 return dtl.DatetimeLikeArrayMixin.astype(self, dtype, copy=copy) ValueError: Cannot convert from timedelta64[ns] to timedelta64[D]. Supported resolutions are 's', 'ms', 'us', 'ns' # to seconds In [65]: td / np.timedelta64(1, 's') Out[65]: 0 2678400.0 1 2678400.0 2 2678703.0 3 NaN dtype: float64 In [66]: td.astype('timedelta64[s]') Out[66]: 0 31 days 00:00:00 1 31 days 00:00:00 2 31 days 00:05:03 3 NaT dtype: timedelta64[s]
Dividing or multiplying a
timedelta64[ns]Series by an integer or integer SeriesIn [67]: td * -1 Out[67]: 0 -31 days +00:00:00 1 -31 days +00:00:00 2 -32 days +23:54:57 3 NaT dtype: timedelta64[ns] In [68]: td * pd.Series([1, 2, 3, 4]) Out[68]: 0 31 days 00:00:00 1 62 days 00:00:00 2 93 days 00:15:09 3 NaT dtype: timedelta64[ns]
Absolute
DateOffsetobjects can act equivalently totimedeltasIn [69]: from pandas import offsets In [70]: td + offsets.Minute(5) + offsets.Milli(5) Out[70]: 0 31 days 00:05:00.005000 1 31 days 00:05:00.005000 2 31 days 00:10:03.005000 3 NaT dtype: timedelta64[ns]
Fillna is now supported for timedeltas
In [71]: td.fillna(pd.Timedelta(0)) Out[71]: 0 31 days 00:00:00 1 31 days 00:00:00 2 31 days 00:05:03 3 0 days 00:00:00 dtype: timedelta64[ns] In [72]: td.fillna(datetime.timedelta(days=1, seconds=5)) Out[72]: 0 31 days 00:00:00 1 31 days 00:00:00 2 31 days 00:05:03 3 1 days 00:00:05 dtype: timedelta64[ns]
You can do numeric reduction operations on timedeltas.
In [73]: td.mean() Out[73]: Timedelta('31 days 00:01:41') In [74]: td.quantile(.1) Out[74]: Timedelta('31 days 00:00:00')
plot(kind='kde')now accepts the optional parametersbw_methodandind, passed to scipy.stats.gaussian_kde() (for scipy >= 0.11.0) to set the bandwidth, and to gkde.evaluate() to specify the indices at which it is evaluated, respectively. See scipy docs. (GH 4298)DataFrame constructor now accepts a numpy masked record array (GH 3478)
The new vectorized string method
extractreturn regular expression matches more conveniently.In [75]: pd.Series(['a1', 'b2', 'c3']).str.extract('[ab](\\d)') Out[75]: 0 0 1 1 2 2 NaN
Elements that do not match return
NaN. Extracting a regular expression with more than one group returns a DataFrame with one column per group.In [76]: pd.Series(['a1', 'b2', 'c3']).str.extract('([ab])(\\d)') Out[76]: 0 1 0 a 1 1 b 2 2 NaN NaN
Elements that do not match return a row of
NaN. Thus, a Series of messy strings can be converted into a like-indexed Series or DataFrame of cleaned-up or more useful strings, without necessitatingget()to access tuples orre.matchobjects.Named groups like
In [77]: pd.Series(['a1', 'b2', 'c3']).str.extract( ....: '(?P<letter>[ab])(?P<digit>\\d)') ....: Out[77]: letter digit 0 a 1 1 b 2 2 NaN NaN
and optional groups can also be used.
In [78]: pd.Series(['a1', 'b2', '3']).str.extract( ....: '(?P<letter>[ab])?(?P<digit>\\d)') ....: Out[78]: letter digit 0 a 1 1 b 2 2 NaN 3
read_statanow accepts Stata 13 format (GH 4291)read_fwfnow infers the column specifications from the first 100 rows of the file if the data has correctly separated and properly aligned columns using the delimiter provided to the function (GH 4488).support for nanosecond times as an offset
Warning
These operations require
numpy >= 1.7Period conversions in the range of seconds and below were reworked and extended up to nanoseconds. Periods in the nanosecond range are now available.
In [79]: pd.date_range('2013-01-01', periods=5, freq='5N') Out[79]: DatetimeIndex([ '2013-01-01 00:00:00', '2013-01-01 00:00:00.000000005', '2013-01-01 00:00:00.000000010', '2013-01-01 00:00:00.000000015', '2013-01-01 00:00:00.000000020'], dtype='datetime64[ns]', freq='5N')
or with frequency as offset
In [80]: pd.date_range('2013-01-01', periods=5, freq=pd.offsets.Nano(5)) Out[80]: DatetimeIndex([ '2013-01-01 00:00:00', '2013-01-01 00:00:00.000000005', '2013-01-01 00:00:00.000000010', '2013-01-01 00:00:00.000000015', '2013-01-01 00:00:00.000000020'], dtype='datetime64[ns]', freq='5N')
Timestamps can be modified in the nanosecond range
In [81]: t = pd.Timestamp('20130101 09:01:02') In [82]: t + pd.tseries.offsets.Nano(123) Out[82]: Timestamp('2013-01-01 09:01:02.000000123')
A new method,
isinfor DataFrames, which plays nicely with boolean indexing. The argument toisin, what we’re comparing the DataFrame to, can be a DataFrame, Series, dict, or array of values. See the docs for more.To get the rows where any of the conditions are met:
In [83]: dfi = pd.DataFrame({'A': [1, 2, 3, 4], 'B': ['a', 'b', 'f', 'n']}) In [84]: dfi Out[84]: A B 0 1 a 1 2 b 2 3 f 3 4 n In [85]: other = pd.DataFrame({'A': [1, 3, 3, 7], 'B': ['e', 'f', 'f', 'e']}) In [86]: mask = dfi.isin(other) In [87]: mask Out[87]: A B 0 True False 1 False False 2 True True 3 False False In [88]: dfi[mask.any(axis=1)] Out[88]: A B 0 1 a 2 3 f
Seriesnow supports ato_framemethod to convert it to a single-column DataFrame (GH 5164)All R datasets listed here http://stat.ethz.ch/R-manual/R-devel/library/datasets/html/00Index.html can now be loaded into pandas objects
# note that pandas.rpy was deprecated in v0.16.0 import pandas.rpy.common as com com.load_data('Titanic')
tz_localizecan infer a fall daylight savings transition based on the structure of the unlocalized data (GH 4230), see the docsDatetimeIndexis now in the API documentation, see the docsjson_normalize()is a new method to allow you to create a flat table from semi-structured JSON data. See the docs (GH 1067)Added PySide support for the qtpandas DataFrameModel and DataFrameWidget.
Python csv parser now supports usecols (GH 4335)
Frequencies gained several new offsets:
DataFrame has a new
interpolatemethod, similar to Series (GH 4434, GH 1892)In [89]: df = pd.DataFrame({'A': [1, 2.1, np.nan, 4.7, 5.6, 6.8], ....: 'B': [.25, np.nan, np.nan, 4, 12.2, 14.4]}) ....: In [90]: df.interpolate() Out[90]: A B 0 1.0 0.25 1 2.1 1.50 2 3.4 2.75 3 4.7 4.00 4 5.6 12.20 5 6.8 14.40
Additionally, the
methodargument tointerpolatehas been expanded to include'nearest', 'zero', 'slinear', 'quadratic', 'cubic', 'barycentric', 'krogh', 'piecewise_polynomial', 'pchip', 'polynomial', 'spline'The new methods require scipy. Consult the Scipy reference guide and documentation for more information about when the various methods are appropriate. See the docs.Interpolate now also accepts a
limitkeyword argument. This works similar tofillna’s limit:In [91]: ser = pd.Series([1, 3, np.nan, np.nan, np.nan, 11]) In [92]: ser.interpolate(limit=2) Out[92]: 0 1.0 1 3.0 2 5.0 3 7.0 4 NaN 5 11.0 dtype: float64
Added
wide_to_longpanel data convenience function. See the docs.In [93]: np.random.seed(123) In [94]: df = pd.DataFrame({"A1970" : {0 : "a", 1 : "b", 2 : "c"}, ....: "A1980" : {0 : "d", 1 : "e", 2 : "f"}, ....: "B1970" : {0 : 2.5, 1 : 1.2, 2 : .7}, ....: "B1980" : {0 : 3.2, 1 : 1.3, 2 : .1}, ....: "X" : dict(zip(range(3), np.random.randn(3))) ....: }) ....: In [95]: df["id"] = df.index In [96]: df Out[96]: A1970 A1980 B1970 B1980 X id 0 a d 2.5 3.2 -1.085631 0 1 b e 1.2 1.3 0.997345 1 2 c f 0.7 0.1 0.282978 2 In [97]: pd.wide_to_long(df, ["A", "B"], i="id", j="year") Out[97]: X A B id year 0 1970 -1.085631 a 2.5 1 1970 0.997345 b 1.2 2 1970 0.282978 c 0.7 0 1980 -1.085631 d 3.2 1 1980 0.997345 e 1.3 2 1980 0.282978 f 0.1
to_csvnow takes adate_formatkeyword argument that specifies how output datetime objects should be formatted. Datetimes encountered in the index, columns, and values will all have this formatting applied. (GH 4313)DataFrame.plotwill scatter plot x versus y by passingkind='scatter'(GH 2215)Added support for Google Analytics v3 API segment IDs that also supports v2 IDs. (GH 5271)
Experimental#
The new
eval()function implements expression evaluation usingnumexprbehind the scenes. This results in large speedups for complicated expressions involving large DataFrames/Series. For example,In [98]: nrows, ncols = 20000, 100 In [99]: df1, df2, df3, df4 = [pd.DataFrame(np.random.randn(nrows, ncols)) ....: for _ in range(4)] ....:
# eval with NumExpr backend In [100]: %timeit pd.eval('df1 + df2 + df3 + df4') 14 ms +- 335 us per loop (mean +- std. dev. of 7 runs, 100 loops each)
# pure Python evaluation In [101]: %timeit df1 + df2 + df3 + df4 12.7 ms +- 280 us per loop (mean +- std. dev. of 7 runs, 100 loops each)
For more details, see the the docs
Similar to
pandas.eval,DataFramehas a newDataFrame.evalmethod that evaluates an expression in the context of theDataFrame. For example,In [102]: df = pd.DataFrame(np.random.randn(10, 2), columns=['a', 'b']) In [103]: df.eval('a + b') Out[103]: 0 -0.685204 1 1.589745 2 0.325441 3 -1.784153 4 -0.432893 5 0.171850 6 1.895919 7 3.065587 8 -0.092759 9 1.391365 dtype: float64
query()method has been added that allows you to select elements of aDataFrameusing a natural query syntax nearly identical to Python syntax. For example,In [104]: n = 20 In [105]: df = pd.DataFrame(np.random.randint(n, size=(n, 3)), columns=['a', 'b', 'c']) In [106]: df.query('a < b < c') Out[106]: a b c 11 1 5 8 15 8 16 19
selects all the rows of
dfwherea < b < cevaluates toTrue. For more details see the the docs.pd.read_msgpack()andpd.to_msgpack()are now a supported method of serialization of arbitrary pandas (and python objects) in a lightweight portable binary format. See the docsWarning
Since this is an EXPERIMENTAL LIBRARY, the storage format may not be stable until a future release.
df = pd.DataFrame(np.random.rand(5, 2), columns=list('AB')) df.to_msgpack('foo.msg') pd.read_msgpack('foo.msg') s = pd.Series(np.random.rand(5), index=pd.date_range('20130101', periods=5)) pd.to_msgpack('foo.msg', df, s) pd.read_msgpack('foo.msg')
You can pass
iterator=Trueto iterator over the unpacked resultsfor o in pd.read_msgpack('foo.msg', iterator=True): print(o)
pandas.io.gbqprovides a simple way to extract from, and load data into, Google’s BigQuery Data Sets by way of pandas DataFrames. BigQuery is a high performance SQL-like database service, useful for performing ad-hoc queries against extremely large datasets. See the docsfrom pandas.io import gbq # A query to select the average monthly temperatures in the # in the year 2000 across the USA. The dataset, # publicata:samples.gsod, is available on all BigQuery accounts, # and is based on NOAA gsod data. query = """SELECT station_number as STATION, month as MONTH, AVG(mean_temp) as MEAN_TEMP FROM publicdata:samples.gsod WHERE YEAR = 2000 GROUP BY STATION, MONTH ORDER BY STATION, MONTH ASC""" # Fetch the result set for this query # Your Google BigQuery Project ID # To find this, see your dashboard: # https://console.developers.google.com/iam-admin/projects?authuser=0 projectid = 'xxxxxxxxx' df = gbq.read_gbq(query, project_id=projectid) # Use pandas to process and reshape the dataset df2 = df.pivot(index='STATION', columns='MONTH', values='MEAN_TEMP') df3 = pd.concat([df2.min(), df2.mean(), df2.max()], axis=1, keys=["Min Tem", "Mean Temp", "Max Temp"])
The resulting DataFrame is:
> df3 Min Tem Mean Temp Max Temp MONTH 1 -53.336667 39.827892 89.770968 2 -49.837500 43.685219 93.437932 3 -77.926087 48.708355 96.099998 4 -82.892858 55.070087 97.317240 5 -92.378261 61.428117 102.042856 6 -77.703334 65.858888 102.900000 7 -87.821428 68.169663 106.510714 8 -89.431999 68.614215 105.500000 9 -86.611112 63.436935 107.142856 10 -78.209677 56.880838 92.103333 11 -50.125000 48.861228 94.996428 12 -50.332258 42.286879 94.396774
Warning
To use this module, you will need a BigQuery account. See <https://cloud.google.com/products/big-query> for details.
As of 10/10/13, there is a bug in Google’s API preventing result sets from being larger than 100,000 rows. A patch is scheduled for the week of 10/14/13.
Internal refactoring#
In 0.13.0 there is a major refactor primarily to subclass Series from
NDFrame, which is the base class currently for DataFrame and Panel,
to unify methods and behaviors. Series formerly subclassed directly from
ndarray. (GH 4080, GH 3862, GH 816)
Warning
There are two potential incompatibilities from < 0.13.0
Using certain numpy functions would previously return a
Seriesif passed aSeriesas an argument. This seems only to affectnp.ones_like,np.empty_like,np.diffandnp.where. These now returnndarrays.In [107]: s = pd.Series([1, 2, 3, 4])
Numpy Usage
In [108]: np.ones_like(s) Out[108]: array([1, 1, 1, 1]) In [109]: np.diff(s) Out[109]: array([1, 1, 1]) In [110]: np.where(s > 1, s, np.nan) Out[110]: array([nan, 2., 3., 4.])
Pandonic Usage
In [111]: pd.Series(1, index=s.index) Out[111]: 0 1 1 1 2 1 3 1 dtype: int64 In [112]: s.diff() Out[112]: 0 NaN 1 1.0 2 1.0 3 1.0 dtype: float64 In [113]: s.where(s > 1) Out[113]: 0 NaN 1 2.0 2 3.0 3 4.0 dtype: float64
Passing a
Seriesdirectly to a cython function expecting anndarraytype will no long work directly, you must passSeries.values, See Enhancing PerformanceSeries(0.5)would previously return the scalar0.5, instead this will return a 1-elementSeriesThis change breaks
rpy2<=2.3.8. an Issue has been opened against rpy2 and a workaround is detailed in GH 5698. Thanks @JanSchulz.
Pickle compatibility is preserved for pickles created prior to 0.13. These must be unpickled with
pd.read_pickle, see Pickling.Refactor of series.py/frame.py/panel.py to move common code to generic.py
added
_setup_axesto created generic NDFrame structuresmoved methods
from_axes,_wrap_array,axes,ix,loc,iloc,shape,empty,swapaxes,transpose,pop__iter__,keys,__contains__,__len__,__neg__,__invert__convert_objects,as_blocks,as_matrix,values__getstate__,__setstate__(compat remains in frame/panel)__getattr__,__setattr___indexed_same,reindex_like,align,where,maskfillna,replace(Seriesreplace is now consistent withDataFrame)filter(also added axis argument to selectively filter on a different axis)reindex,reindex_axis,taketruncate(moved to become part ofNDFrame)
These are API changes which make
Panelmore consistent withDataFrameswapaxeson aPanelwith the same axes specified now return a copysupport attribute access for setting
filter supports the same API as the original
DataFramefilter
Reindex called with no arguments will now return a copy of the input object
TimeSeriesis now an alias forSeries. the propertyis_time_seriescan be used to distinguish (if desired)Refactor of Sparse objects to use BlockManager
Created a new block type in internals,
SparseBlock, which can hold multi-dtypes and is non-consolidatable.SparseSeriesandSparseDataFramenow inherit more methods from there hierarchy (Series/DataFrame), and no longer inherit fromSparseArray(which instead is the object of theSparseBlock)Sparse suite now supports integration with non-sparse data. Non-float sparse data is supportable (partially implemented)
Operations on sparse structures within DataFrames should preserve sparseness, merging type operations will convert to dense (and back to sparse), so might be somewhat inefficient
enable setitem on
SparseSeriesfor boolean/integer/slicesSparsePanelsimplementation is unchanged (e.g. not using BlockManager, needs work)
added
ftypesmethod to Series/DataFrame, similar todtypes, but indicates if the underlying is sparse/dense (as well as the dtype)All
NDFrameobjects can now use__finalize__()to specify various values to propagate to new objects from an existing one (e.g.nameinSerieswill follow more automatically now)Internal type checking is now done via a suite of generated classes, allowing
isinstance(value, klass)without having to directly import the klass, courtesy of @jtratnerBug in Series update where the parent frame is not updating its cache based on changes (GH 4080) or types (GH 3217), fillna (GH 3386)
Refactor
Series.reindexto core/generic.py (GH 4604, GH 4618), allowmethod=in reindexing on a Series to workSeries.copyno longer accepts theorderparameter and is now consistent withNDFramecopyRefactor
renamemethods to core/generic.py; fixesSeries.renamefor (GH 4605), and addsrenamewith the same signature forPanelRefactor
clipmethods to core/generic.py (GH 4798)Refactor of
_get_numeric_data/_get_bool_datato core/generic.py, allowing Series/Panel functionalitySeries(for index) /Panel(for items) now allow attribute access to its elements (GH 1903)In [114]: s = pd.Series([1, 2, 3], index=list('abc')) In [115]: s.b Out[115]: 2 In [116]: s.a = 5 In [117]: s Out[117]: a 5 b 2 c 3 dtype: int64
Bug fixes#
HDFStoreraising an invalid
TypeErrorrather thanValueErrorwhen appending with a different block ordering (GH 4096)read_hdfwas not respecting as passedmode(GH 4504)appending a 0-len table will work correctly (GH 4273)
to_hdfwas raising when passing both argumentsappendandtable(GH 4584)reading from a store with duplicate columns across dtypes would raise (GH 4767)
Fixed a bug where
ValueErrorwasn’t correctly raised when column names weren’t strings (GH 4956)A zero length series written in Fixed format not deserializing properly. (GH 4708)
Fixed decoding perf issue on pyt3 (GH 5441)
Validate levels in a MultiIndex before storing (GH 5527)
Correctly handle
data_columnswith a Panel (GH 5717)
Fixed bug in tslib.tz_convert(vals, tz1, tz2): it could raise IndexError exception while trying to access trans[pos + 1] (GH 4496)
The
byargument now works correctly with thelayoutargument (GH 4102, GH 4014) in*.histplotting methodsFixed bug in
PeriodIndex.mapwhere usingstrwould return the str representation of the index (GH 4136)Fixed test failure
test_time_series_plot_color_with_empty_kwargswhen using custom matplotlib default colors (GH 4345)Fix running of stata IO tests. Now uses temporary files to write (GH 4353)
Fixed an issue where
DataFrame.sumwas slower thanDataFrame.meanfor integer valued frames (GH 4365)read_htmltests now work with Python 2.6 (GH 4351)Fixed bug where
networktesting was throwingNameErrorbecause a local variable was undefined (GH 4381)In
to_json, raise if a passedorientwould cause loss of data because of a duplicate index (GH 4359)In
to_json, fix date handling so milliseconds are the default timestamp as the docstring says (GH 4362).as_indexis no longer ignored when doing groupby apply (GH 4648, GH 3417)JSON NaT handling fixed, NaTs are now serialized to
null(GH 4498)Fixed JSON handling of escapable characters in JSON object keys (GH 4593)
Fixed passing
keep_default_na=Falsewhenna_values=None(GH 4318)Fixed bug with
valuesraising an error on a DataFrame with duplicate columns and mixed dtypes, surfaced in (GH 4377)Fixed bug with duplicate columns and type conversion in
read_jsonwhenorient='split'(GH 4377)Fixed JSON bug where locales with decimal separators other than ‘.’ threw exceptions when encoding / decoding certain values. (GH 4918)
Fix
.iatindexing with aPeriodIndex(GH 4390)Fixed an issue where
PeriodIndexjoining with self was returning a new instance rather than the same instance (GH 4379); also adds a test for this for the other index typesFixed a bug with all the dtypes being converted to object when using the CSV cparser with the usecols parameter (GH 3192)
Fix an issue in merging blocks where the resulting DataFrame had partially set _ref_locs (GH 4403)
Fixed an issue where hist subplots were being overwritten when they were called using the top level matplotlib API (GH 4408)
Fixed a bug where calling
Series.astype(str)would truncate the string (GH 4405, GH 4437)Fixed a py3 compat issue where bytes were being repr’d as tuples (GH 4455)
Fixed Panel attribute naming conflict if item is named ‘a’ (GH 3440)
Fixed an issue where duplicate indexes were raising when plotting (GH 4486)
Fixed an issue where cumsum and cumprod didn’t work with bool dtypes (GH 4170, GH 4440)
Fixed Panel slicing issued in
xsthat was returning an incorrect dimmed object (GH 4016)Fix resampling bug where custom reduce function not used if only one group (GH 3849, GH 4494)
Fixed Panel assignment with a transposed frame (GH 3830)
Raise on set indexing with a Panel and a Panel as a value which needs alignment (GH 3777)
frozenset objects now raise in the
Seriesconstructor (GH 4482, GH 4480)Fixed issue with sorting a duplicate MultiIndex that has multiple dtypes (GH 4516)
Fixed bug in
DataFrame.set_valueswhich was causing name attributes to be lost when expanding the index. (GH 3742, GH 4039)Fixed issue where individual
names,levelsandlabelscould be set onMultiIndexwithout validation (GH 3714, GH 4039)Fixed (GH 3334) in pivot_table. Margins did not compute if values is the index.
Fix bug in having a rhs of
np.timedelta64ornp.offsets.DateOffsetwhen operating with datetimes (GH 4532)Fix arithmetic with series/datetimeindex and
np.timedelta64not working the same (GH 4134) and buggy timedelta in NumPy 1.6 (GH 4135)Fix bug in
pd.read_clipboardon windows with PY3 (GH 4561); not decoding properlytslib.get_period_field()andtslib.get_period_field_arr()now raise if code argument out of range (GH 4519, GH 4520)Fix boolean indexing on an empty series loses index names (GH 4235), infer_dtype works with empty arrays.
Fix reindexing with multiple axes; if an axes match was not replacing the current axes, leading to a possible lazy frequency inference issue (GH 3317)
Fixed issue where
DataFrame.applywas reraising exceptions incorrectly (causing the original stack trace to be truncated).Fix selection with
ix/locand non_unique selectors (GH 4619)Fix assignment with iloc/loc involving a dtype change in an existing column (GH 4312, GH 5702) have internal setitem_with_indexer in core/indexing to use Block.setitem
Fixed bug where thousands operator was not handled correctly for floating point numbers in csv_import (GH 4322)
Fix an issue with CacheableOffset not properly being used by many DateOffset; this prevented the DateOffset from being cached (GH 4609)
Fix boolean comparison with a DataFrame on the lhs, and a list/tuple on the rhs (GH 4576)
Fix error/dtype conversion with setitem of
NoneonSeries/DataFrame(GH 4667)Fix decoding based on a passed in non-default encoding in
pd.read_stata(GH 4626)Fix
DataFrame.from_recordswith a plain-vanillandarray. (GH 4727)Fix some inconsistencies with
Index.renameandMultiIndex.rename, etc. (GH 4718, GH 4628)Bug in using
iloc/locwith a cross-sectional and duplicate indices (GH 4726)Bug with using
QUOTE_NONEwithto_csvcausingException. (GH 4328)Bug with Series indexing not raising an error when the right-hand-side has an incorrect length (GH 2702)
Bug in MultiIndexing with a partial string selection as one part of a MultIndex (GH 4758)
Bug with reindexing on the index with a non-unique index will now raise
ValueError(GH 4746)Bug in setting with
loc/ixa single indexer with a MultiIndex axis and a NumPy array, related to (GH 3777)Bug in concatenation with duplicate columns across dtypes not merging with axis=0 (GH 4771, GH 4975)
Bug in
ilocwith a slice index failing (GH 4771)Incorrect error message with no colspecs or width in
read_fwf. (GH 4774)Fix bugs in indexing in a Series with a duplicate index (GH 4548, GH 4550)
Fixed bug with reading compressed files with
read_fwfin Python 3. (GH 3963)Fixed an issue with a duplicate index and assignment with a dtype change (GH 4686)
Fixed bug with reading compressed files in as
bytesrather thanstrin Python 3. Simplifies bytes-producing file-handling in Python 3 (GH 3963, GH 4785).Fixed an issue related to ticklocs/ticklabels with log scale bar plots across different versions of matplotlib (GH 4789)
Suppressed DeprecationWarning associated with internal calls issued by repr() (GH 4391)
Fixed an issue with a duplicate index and duplicate selector with
.loc(GH 4825)Fixed an issue with
DataFrame.sort_indexwhere, when sorting by a single column and passing a list forascending, the argument forascendingwas being interpreted asTrue(GH 4839, GH 4846)Fixed
Panel.tshiftnot working. Addedfreqsupport toPanel.shift(GH 4853)Fix an issue in TextFileReader w/ Python engine (i.e. PythonParser) with thousands != “,” (GH 4596)
Bug in getitem with a duplicate index when using where (GH 4879)
Fix Type inference code coerces float column into datetime (GH 4601)
Fixed
_ensure_numericdoes not check for complex numbers (GH 4902)Fixed a bug in
Series.histwhere two figures were being created when thebyargument was passed (GH 4112, GH 4113).Fixed a bug in
convert_objectsfor > 2 ndims (GH 4937)Fixed a bug in DataFrame/Panel cache insertion and subsequent indexing (GH 4939, GH 5424)
Fixed string methods for
FrozenNDArrayandFrozenList(GH 4929)Fixed a bug with setting invalid or out-of-range values in indexing enlargement scenarios (GH 4940)
Tests for fillna on empty Series (GH 4346), thanks @immerrr
Fixed
copy()to shallow copy axes/indices as well and thereby keep separate metadata. (GH 4202, GH 4830)Fixed skiprows option in Python parser for read_csv (GH 4382)
Fixed bug preventing
cutfrom working withnp.inflevels without explicitly passing labels (GH 3415)Fixed wrong check for overlapping in
DatetimeIndex.union(GH 4564)Fixed conflict between thousands separator and date parser in csv_parser (GH 4678)
Fix appending when dtypes are not the same (error showing mixing float/np.datetime64) (GH 4993)
Fix repr for DateOffset. No longer show duplicate entries in kwds. Removed unused offset fields. (GH 4638)
Fixed wrong index name during read_csv if using usecols. Applies to c parser only. (GH 4201)
Timestampobjects can now appear in the left hand side of a comparison operation with aSeriesorDataFrameobject (GH 4982).Fix a bug when indexing with
np.nanviailoc/loc(GH 5016)Fixed a bug where low memory c parser could create different types in different chunks of the same file. Now coerces to numerical type or raises warning. (GH 3866)
Fix a bug where reshaping a
Seriesto its own shape raisedTypeError(GH 4554) and other reshaping issues.Bug in setting with
ix/locand a mixed int/string index (GH 4544)Make sure series-series boolean comparisons are label based (GH 4947)
Bug in multi-level indexing with a Timestamp partial indexer (GH 4294)
Tests/fix for MultiIndex construction of an all-nan frame (GH 4078)
Fixed a bug where
read_html()wasn’t correctly inferring values of tables with commas (GH 5029)Fixed a bug where
read_html()wasn’t providing a stable ordering of returned tables (GH 4770, GH 5029).Fixed a bug where
read_html()was incorrectly parsing when passedindex_col=0(GH 5066).Fixed a bug where
read_html()was incorrectly inferring the type of headers (GH 5048).Fixed a bug where
DatetimeIndexjoins withPeriodIndexcaused a stack overflow (GH 3899).Fixed a bug where
groupbyobjects didn’t allow plots (GH 5102).Fixed a bug where
groupbyobjects weren’t tab-completing column names (GH 5102).Fixed a bug where
groupby.plot()and friends were duplicating figures multiple times (GH 5102).Provide automatic conversion of
objectdtypes on fillna, related (GH 5103)Fixed a bug where default options were being overwritten in the option parser cleaning (GH 5121).
Treat a list/ndarray identically for
ilocindexing with list-like (GH 5006)Fix
MultiIndex.get_level_values()with missing values (GH 5074)Fix bound checking for Timestamp() with datetime64 input (GH 4065)
Fix a bug where
TestReadHtmlwasn’t calling the correctread_html()function (GH 5150).Fix a bug with
NDFrame.replace()which made replacement appear as though it was (incorrectly) using regular expressions (GH 5143).Fix better error message for to_datetime (GH 4928)
Made sure different locales are tested on travis-ci (GH 4918). Also adds a couple of utilities for getting locales and setting locales with a context manager.
Fixed segfault on
isnull(MultiIndex)(now raises an error instead) (GH 5123, GH 5125)Allow duplicate indices when performing operations that align (GH 5185, GH 5639)
Compound dtypes in a constructor raise
NotImplementedError(GH 5191)Bug in comparing duplicate frames (GH 4421) related
Bug in describe on duplicate frames
Bug in
to_datetimewith a format andcoerce=Truenot raising (GH 5195)Bug in
locsetting with multiple indexers and a rhs of a Series that needs broadcasting (GH 5206)Fixed bug where inplace setting of levels or labels on
MultiIndexwould not clear cachedvaluesproperty and therefore return wrongvalues. (GH 5215)Fixed bug where filtering a grouped DataFrame or Series did not maintain the original ordering (GH 4621).
Fixed
Periodwith a business date freq to always roll-forward if on a non-business date. (GH 5203)Fixed bug in Excel writers where frames with duplicate column names weren’t written correctly. (GH 5235)
Fixed issue with
dropand a non-unique index on Series (GH 5248)Fixed segfault in C parser caused by passing more names than columns in the file. (GH 5156)
Fix
Series.isinwith date/time-like dtypes (GH 5021)C and Python Parser can now handle the more common MultiIndex column format which doesn’t have a row for index names (GH 4702)
Bug when trying to use an out-of-bounds date as an object dtype (GH 5312)
Bug when trying to display an embedded PandasObject (GH 5324)
Allows operating of Timestamps to return a datetime if the result is out-of-bounds related (GH 5312)
Fix return value/type signature of
initObjToJSON()to be compatible with numpy’simport_array()(GH 5334, GH 5326)Bug when renaming then set_index on a DataFrame (GH 5344)
Test suite no longer leaves around temporary files when testing graphics. (GH 5347) (thanks for catching this @yarikoptic!)
Fixed html tests on win32. (GH 4580)
Make sure that
head/tailareilocbased, (GH 5370)Fixed bug for
PeriodIndexstring representation if there are 1 or 2 elements. (GH 5372)The GroupBy methods
transformandfiltercan be used on Series and DataFrames that have repeated (non-unique) indices. (GH 4620)Fix empty series not printing name in repr (GH 4651)
Make tests create temp files in temp directory by default. (GH 5419)
pd.to_timedeltaof a scalar returns a scalar (GH 5410)pd.to_timedeltaacceptsNaNandNaT, returningNaTinstead of raising (GH 5437)performance improvements in
isnullon larger size pandas objectsFixed various setitem with 1d ndarray that does not have a matching length to the indexer (GH 5508)
Bug in getitem with a MultiIndex and
iloc(GH 5528)Bug in delitem on a Series (GH 5542)
Bug fix in apply when using custom function and objects are not mutated (GH 5545)
Bug in selecting from a non-unique index with
loc(GH 5553)Bug in groupby returning non-consistent types when user function returns a
None, (GH 5592)Work around regression in numpy 1.7.0 which erroneously raises IndexError from
ndarray.item(GH 5666)Bug in repeated indexing of object with resultant non-unique index (GH 5678)
Bug in fillna with Series and a passed series/dict (GH 5703)
Bug in groupby transform with a datetime-like grouper (GH 5712)
Bug in MultiIndex selection in PY3 when using certain keys (GH 5725)
Row-wise concat of differing dtypes failing in certain cases (GH 5754)
Contributors#
A total of 77 people contributed patches to this release. People with a “+” by their names contributed a patch for the first time.
Agustín Herranz +
Alex Gaudio +
Alex Rothberg +
Andreas Klostermann +
Andreas Würl +
Andy Hayden
Ben Alex +
Benedikt Sauer +
Brad Buran
Caleb Epstein +
Chang She
Christopher Whelan
DSM +
Dale Jung +
Dan Birken
David Rasch +
Dieter Vandenbussche
Gabi Davar +
Garrett Drapala
Goyo +
Greg Reda +
Ivan Smirnov +
Jack Kelly +
Jacob Schaer +
Jan Schulz +
Jeff Tratner
Jeffrey Tratner
John McNamara +
John W. O’Brien +
Joris Van den Bossche
Justin Bozonier +
Kelsey Jordahl
Kevin Stone
Kieran O’Mahony
Kyle Hausmann +
Kyle Kelley +
Kyle Meyer
Mike Kelly
Mortada Mehyar +
Nick Foti +
Olivier Harris +
Ondřej Čertík +
PKEuS
Phillip Cloud
Pierre Haessig +
Richard T. Guy +
Roman Pekar +
Roy Hyunjin Han
Skipper Seabold
Sten +
Thomas A Caswell +
Thomas Kluyver
Tiago Requeijo +
TomAugspurger
Trent Hauck
Valentin Haenel +
Viktor Kerkez +
Vincent Arel-Bundock
Wes McKinney
Wes Turner +
Weston Renoud +
Yaroslav Halchenko
Zach Dwiel +
chapman siu +
chappers +
d10genes +
danielballan
daydreamt +
engstrom +
jreback
monicaBee +
prossahl +
rockg +
unutbu +
westurner +
y-p
zach powers