10 minutes to pandas#
This is a short introduction to pandas, geared mainly for new users. You can see more complex recipes in the Cookbook.
Customarily, we import as follows:
In [1]: import numpy as np
In [2]: import pandas as pd
Object creation#
See the Intro to data structures section.
Creating a Series by passing a list of values, letting pandas create
a default integer index:
In [3]: s = pd.Series([1, 3, 5, np.nan, 6, 8])
In [4]: s
Out[4]:
0 1.0
1 3.0
2 5.0
3 NaN
4 6.0
5 8.0
dtype: float64
Creating a DataFrame by passing a NumPy array, with a datetime index using date_range()
and labeled columns:
In [5]: dates = pd.date_range("20130101", periods=6)
In [6]: dates
Out[6]:
DatetimeIndex(['2013-01-01', '2013-01-02', '2013-01-03', '2013-01-04',
'2013-01-05', '2013-01-06'],
dtype='datetime64[ns]', freq='D')
In [7]: df = pd.DataFrame(np.random.randn(6, 4), index=dates, columns=list("ABCD"))
In [8]: df
Out[8]:
A B C D
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
2013-01-05 -0.424972 0.567020 0.276232 -1.087401
2013-01-06 -0.673690 0.113648 -1.478427 0.524988
Creating a DataFrame by passing a dictionary of objects that can be
converted into a series-like structure:
In [9]: df2 = pd.DataFrame(
...: {
...: "A": 1.0,
...: "B": pd.Timestamp("20130102"),
...: "C": pd.Series(1, index=list(range(4)), dtype="float32"),
...: "D": np.array([3] * 4, dtype="int32"),
...: "E": pd.Categorical(["test", "train", "test", "train"]),
...: "F": "foo",
...: }
...: )
...:
In [10]: df2
Out[10]:
A B C D E F
0 1.0 2013-01-02 1.0 3 test foo
1 1.0 2013-01-02 1.0 3 train foo
2 1.0 2013-01-02 1.0 3 test foo
3 1.0 2013-01-02 1.0 3 train foo
The columns of the resulting DataFrame have different
dtypes:
In [11]: df2.dtypes
Out[11]:
A float64
B datetime64[ns]
C float32
D int32
E category
F object
dtype: object
If you’re using IPython, tab completion for column names (as well as public attributes) is automatically enabled. Here’s a subset of the attributes that will be completed:
In [12]: df2.<TAB> # noqa: E225, E999
df2.A df2.bool
df2.abs df2.boxplot
df2.add df2.C
df2.add_prefix df2.clip
df2.add_suffix df2.columns
df2.align df2.copy
df2.all df2.count
df2.any df2.combine
df2.append df2.D
df2.apply df2.describe
df2.applymap df2.diff
df2.B df2.duplicated
As you can see, the columns A, B, C, and D are automatically
tab completed. E and F are there as well; the rest of the attributes have been
truncated for brevity.
Viewing data#
See the Basics section.
Use DataFrame.head() and DataFrame.tail() to view the top and bottom rows of the frame
respectively:
In [13]: df.head()
Out[13]:
A B C D
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
2013-01-05 -0.424972 0.567020 0.276232 -1.087401
In [14]: df.tail(3)
Out[14]:
A B C D
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
2013-01-05 -0.424972 0.567020 0.276232 -1.087401
2013-01-06 -0.673690 0.113648 -1.478427 0.524988
Display the DataFrame.index or DataFrame.columns:
In [15]: df.index
Out[15]:
DatetimeIndex(['2013-01-01', '2013-01-02', '2013-01-03', '2013-01-04',
'2013-01-05', '2013-01-06'],
dtype='datetime64[ns]', freq='D')
In [16]: df.columns
Out[16]: Index(['A', 'B', 'C', 'D'], dtype='object')
DataFrame.to_numpy() gives a NumPy representation of the underlying data.
Note that this can be an expensive operation when your DataFrame has
columns with different data types, which comes down to a fundamental difference
between pandas and NumPy: NumPy arrays have one dtype for the entire array,
while pandas DataFrames have one dtype per column. When you call
DataFrame.to_numpy(), pandas will find the NumPy dtype that can hold all
of the dtypes in the DataFrame. This may end up being object, which requires
casting every value to a Python object.
For df, our DataFrame of all floating-point values, and
DataFrame.to_numpy() is fast and doesn’t require copying data:
In [17]: df.to_numpy()
Out[17]:
array([[ 0.4691, -0.2829, -1.5091, -1.1356],
[ 1.2121, -0.1732, 0.1192, -1.0442],
[-0.8618, -2.1046, -0.4949, 1.0718],
[ 0.7216, -0.7068, -1.0396, 0.2719],
[-0.425 , 0.567 , 0.2762, -1.0874],
[-0.6737, 0.1136, -1.4784, 0.525 ]])
For df2, the DataFrame with multiple dtypes,
DataFrame.to_numpy() is relatively expensive:
In [18]: df2.to_numpy()
Out[18]:
array([[1.0, Timestamp('2013-01-02 00:00:00'), 1.0, 3, 'test', 'foo'],
[1.0, Timestamp('2013-01-02 00:00:00'), 1.0, 3, 'train', 'foo'],
[1.0, Timestamp('2013-01-02 00:00:00'), 1.0, 3, 'test', 'foo'],
[1.0, Timestamp('2013-01-02 00:00:00'), 1.0, 3, 'train', 'foo']],
dtype=object)
Note
DataFrame.to_numpy() does not include the index or column
labels in the output.
describe() shows a quick statistic summary of your data:
In [19]: df.describe()
Out[19]:
A B C D
count 6.000000 6.000000 6.000000 6.000000
mean 0.073711 -0.431125 -0.687758 -0.233103
std 0.843157 0.922818 0.779887 0.973118
min -0.861849 -2.104569 -1.509059 -1.135632
25% -0.611510 -0.600794 -1.368714 -1.076610
50% 0.022070 -0.228039 -0.767252 -0.386188
75% 0.658444 0.041933 -0.034326 0.461706
max 1.212112 0.567020 0.276232 1.071804
Transposing your data:
In [20]: df.T
Out[20]:
2013-01-01 2013-01-02 2013-01-03 2013-01-04 2013-01-05 2013-01-06
A 0.469112 1.212112 -0.861849 0.721555 -0.424972 -0.673690
B -0.282863 -0.173215 -2.104569 -0.706771 0.567020 0.113648
C -1.509059 0.119209 -0.494929 -1.039575 0.276232 -1.478427
D -1.135632 -1.044236 1.071804 0.271860 -1.087401 0.524988
DataFrame.sort_index() sorts by an axis:
In [21]: df.sort_index(axis=1, ascending=False)
Out[21]:
D C B A
2013-01-01 -1.135632 -1.509059 -0.282863 0.469112
2013-01-02 -1.044236 0.119209 -0.173215 1.212112
2013-01-03 1.071804 -0.494929 -2.104569 -0.861849
2013-01-04 0.271860 -1.039575 -0.706771 0.721555
2013-01-05 -1.087401 0.276232 0.567020 -0.424972
2013-01-06 0.524988 -1.478427 0.113648 -0.673690
DataFrame.sort_values() sorts by values:
In [22]: df.sort_values(by="B")
Out[22]:
A B C D
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-06 -0.673690 0.113648 -1.478427 0.524988
2013-01-05 -0.424972 0.567020 0.276232 -1.087401
Selection#
Note
While standard Python / NumPy expressions for selecting and setting are
intuitive and come in handy for interactive work, for production code, we
recommend the optimized pandas data access methods, DataFrame.at(), DataFrame.iat(),
DataFrame.loc() and DataFrame.iloc().
See the indexing documentation Indexing and Selecting Data and MultiIndex / Advanced Indexing.
Getting#
Selecting a single column, which yields a Series,
equivalent to df.A:
In [23]: df["A"]
Out[23]:
2013-01-01 0.469112
2013-01-02 1.212112
2013-01-03 -0.861849
2013-01-04 0.721555
2013-01-05 -0.424972
2013-01-06 -0.673690
Freq: D, Name: A, dtype: float64
Selecting via [] (__getitem__), which slices the rows:
In [24]: df[0:3]
Out[24]:
A B C D
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
In [25]: df["20130102":"20130104"]
Out[25]:
A B C D
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
Selection by label#
See more in Selection by Label using DataFrame.loc() or DataFrame.at().
For getting a cross section using a label:
In [26]: df.loc[dates[0]]
Out[26]:
A 0.469112
B -0.282863
C -1.509059
D -1.135632
Name: 2013-01-01 00:00:00, dtype: float64
Selecting on a multi-axis by label:
In [27]: df.loc[:, ["A", "B"]]
Out[27]:
A B
2013-01-01 0.469112 -0.282863
2013-01-02 1.212112 -0.173215
2013-01-03 -0.861849 -2.104569
2013-01-04 0.721555 -0.706771
2013-01-05 -0.424972 0.567020
2013-01-06 -0.673690 0.113648
Showing label slicing, both endpoints are included:
In [28]: df.loc["20130102":"20130104", ["A", "B"]]
Out[28]:
A B
2013-01-02 1.212112 -0.173215
2013-01-03 -0.861849 -2.104569
2013-01-04 0.721555 -0.706771
Reduction in the dimensions of the returned object:
In [29]: df.loc["20130102", ["A", "B"]]
Out[29]:
A 1.212112
B -0.173215
Name: 2013-01-02 00:00:00, dtype: float64
For getting a scalar value:
In [30]: df.loc[dates[0], "A"]
Out[30]: 0.4691122999071863
For getting fast access to a scalar (equivalent to the prior method):
In [31]: df.at[dates[0], "A"]
Out[31]: 0.4691122999071863
Selection by position#
See more in Selection by Position using DataFrame.iloc() or DataFrame.at().
Select via the position of the passed integers:
In [32]: df.iloc[3]
Out[32]:
A 0.721555
B -0.706771
C -1.039575
D 0.271860
Name: 2013-01-04 00:00:00, dtype: float64
By integer slices, acting similar to NumPy/Python:
In [33]: df.iloc[3:5, 0:2]
Out[33]:
A B
2013-01-04 0.721555 -0.706771
2013-01-05 -0.424972 0.567020
By lists of integer position locations, similar to the NumPy/Python style:
In [34]: df.iloc[[1, 2, 4], [0, 2]]
Out[34]:
A C
2013-01-02 1.212112 0.119209
2013-01-03 -0.861849 -0.494929
2013-01-05 -0.424972 0.276232
For slicing rows explicitly:
In [35]: df.iloc[1:3, :]
Out[35]:
A B C D
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804
For slicing columns explicitly:
In [36]: df.iloc[:, 1:3]
Out[36]:
B C
2013-01-01 -0.282863 -1.509059
2013-01-02 -0.173215 0.119209
2013-01-03 -2.104569 -0.494929
2013-01-04 -0.706771 -1.039575
2013-01-05 0.567020 0.276232
2013-01-06 0.113648 -1.478427
For getting a value explicitly:
In [37]: df.iloc[1, 1]
Out[37]: -0.17321464905330858
For getting fast access to a scalar (equivalent to the prior method):
In [38]: df.iat[1, 1]
Out[38]: -0.17321464905330858
Boolean indexing#
Using a single column’s values to select data:
In [39]: df[df["A"] > 0]
Out[39]:
A B C D
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632
2013-01-02 1.212112 -0.173215 0.119209 -1.044236
2013-01-04 0.721555 -0.706771 -1.039575 0.271860
Selecting values from a DataFrame where a boolean condition is met:
In [40]: df[df > 0]
Out[40]:
A B C D
2013-01-01 0.469112 NaN NaN NaN
2013-01-02 1.212112 NaN 0.119209 NaN
2013-01-03 NaN NaN NaN 1.071804
2013-01-04 0.721555 NaN NaN 0.271860
2013-01-05 NaN 0.567020 0.276232 NaN
2013-01-06 NaN 0.113648 NaN 0.524988
Using the isin() method for filtering:
In [41]: df2 = df.copy()
In [42]: df2["E"] = ["one", "one", "two", "three", "four", "three"]
In [43]: df2
Out[43]:
A B C D E
2013-01-01 0.469112 -0.282863 -1.509059 -1.135632 one
2013-01-02 1.212112 -0.173215 0.119209 -1.044236 one
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804 two
2013-01-04 0.721555 -0.706771 -1.039575 0.271860 three
2013-01-05 -0.424972 0.567020 0.276232 -1.087401 four
2013-01-06 -0.673690 0.113648 -1.478427 0.524988 three
In [44]: df2[df2["E"].isin(["two", "four"])]
Out[44]:
A B C D E
2013-01-03 -0.861849 -2.104569 -0.494929 1.071804 two
2013-01-05 -0.424972 0.567020 0.276232 -1.087401 four
Setting#
Setting a new column automatically aligns the data by the indexes:
In [45]: s1 = pd.Series([1, 2, 3, 4, 5, 6], index=pd.date_range("20130102", periods=6))
In [46]: s1
Out[46]:
2013-01-02 1
2013-01-03 2
2013-01-04 3
2013-01-05 4
2013-01-06 5
2013-01-07 6
Freq: D, dtype: int64
In [47]: df["F"] = s1
Setting values by label:
In [48]: df.at[dates[0], "A"] = 0
Setting values by position:
In [49]: df.iat[0, 1] = 0
Setting by assigning with a NumPy array:
In [50]: df.loc[:, "D"] = np.array([5] * len(df))
The result of the prior setting operations:
In [51]: df
Out[51]:
A B C D F
2013-01-01 0.000000 0.000000 -1.509059 5 NaN
2013-01-02 1.212112 -0.173215 0.119209 5 1.0
2013-01-03 -0.861849 -2.104569 -0.494929 5 2.0
2013-01-04 0.721555 -0.706771 -1.039575 5 3.0
2013-01-05 -0.424972 0.567020 0.276232 5 4.0
2013-01-06 -0.673690 0.113648 -1.478427 5 5.0
A where operation with setting:
In [52]: df2 = df.copy()
In [53]: df2[df2 > 0] = -df2
In [54]: df2
Out[54]:
A B C D F
2013-01-01 0.000000 0.000000 -1.509059 -5 NaN
2013-01-02 -1.212112 -0.173215 -0.119209 -5 -1.0
2013-01-03 -0.861849 -2.104569 -0.494929 -5 -2.0
2013-01-04 -0.721555 -0.706771 -1.039575 -5 -3.0
2013-01-05 -0.424972 -0.567020 -0.276232 -5 -4.0
2013-01-06 -0.673690 -0.113648 -1.478427 -5 -5.0
Missing data#
pandas primarily uses the value np.nan to represent missing data. It is by
default not included in computations. See the Missing Data section.
Reindexing allows you to change/add/delete the index on a specified axis. This returns a copy of the data:
In [55]: df1 = df.reindex(index=dates[0:4], columns=list(df.columns) + ["E"])
In [56]: df1.loc[dates[0] : dates[1], "E"] = 1
In [57]: df1
Out[57]:
A B C D F E
2013-01-01 0.000000 0.000000 -1.509059 5 NaN 1.0
2013-01-02 1.212112 -0.173215 0.119209 5 1.0 1.0
2013-01-03 -0.861849 -2.104569 -0.494929 5 2.0 NaN
2013-01-04 0.721555 -0.706771 -1.039575 5 3.0 NaN
DataFrame.dropna() drops any rows that have missing data:
In [58]: df1.dropna(how="any")
Out[58]:
A B C D F E
2013-01-02 1.212112 -0.173215 0.119209 5 1.0 1.0
DataFrame.fillna() fills missing data:
In [59]: df1.fillna(value=5)
Out[59]:
A B C D F E
2013-01-01 0.000000 0.000000 -1.509059 5 5.0 1.0
2013-01-02 1.212112 -0.173215 0.119209 5 1.0 1.0
2013-01-03 -0.861849 -2.104569 -0.494929 5 2.0 5.0
2013-01-04 0.721555 -0.706771 -1.039575 5 3.0 5.0
isna() gets the boolean mask where values are nan:
In [60]: pd.isna(df1)
Out[60]:
A B C D F E
2013-01-01 False False False False True False
2013-01-02 False False False False False False
2013-01-03 False False False False False True
2013-01-04 False False False False False True
Operations#
See the Basic section on Binary Ops.
Stats#
Operations in general exclude missing data.
Performing a descriptive statistic:
In [61]: df.mean()
Out[61]:
A -0.004474
B -0.383981
C -0.687758
D 5.000000
F 3.000000
dtype: float64
Same operation on the other axis:
In [62]: df.mean(1)
Out[62]:
2013-01-01 0.872735
2013-01-02 1.431621
2013-01-03 0.707731
2013-01-04 1.395042
2013-01-05 1.883656
2013-01-06 1.592306
Freq: D, dtype: float64
Operating with objects that have different dimensionality and need alignment. In addition, pandas automatically broadcasts along the specified dimension:
In [63]: s = pd.Series([1, 3, 5, np.nan, 6, 8], index=dates).shift(2)
In [64]: s
Out[64]:
2013-01-01 NaN
2013-01-02 NaN
2013-01-03 1.0
2013-01-04 3.0
2013-01-05 5.0
2013-01-06 NaN
Freq: D, dtype: float64
In [65]: df.sub(s, axis="index")
Out[65]:
A B C D F
2013-01-01 NaN NaN NaN NaN NaN
2013-01-02 NaN NaN NaN NaN NaN
2013-01-03 -1.861849 -3.104569 -1.494929 4.0 1.0
2013-01-04 -2.278445 -3.706771 -4.039575 2.0 0.0
2013-01-05 -5.424972 -4.432980 -4.723768 0.0 -1.0
2013-01-06 NaN NaN NaN NaN NaN
Apply#
DataFrame.apply() applies a user defined function to the data:
In [66]: df.apply(np.cumsum)
Out[66]:
A B C D F
2013-01-01 0.000000 0.000000 -1.509059 5 NaN
2013-01-02 1.212112 -0.173215 -1.389850 10 1.0
2013-01-03 0.350263 -2.277784 -1.884779 15 3.0
2013-01-04 1.071818 -2.984555 -2.924354 20 6.0
2013-01-05 0.646846 -2.417535 -2.648122 25 10.0
2013-01-06 -0.026844 -2.303886 -4.126549 30 15.0
In [67]: df.apply(lambda x: x.max() - x.min())
Out[67]:
A 2.073961
B 2.671590
C 1.785291
D 0.000000
F 4.000000
dtype: float64
Histogramming#
See more at Histogramming and Discretization.
In [68]: s = pd.Series(np.random.randint(0, 7, size=10))
In [69]: s
Out[69]:
0 4
1 2
2 1
3 2
4 6
5 4
6 4
7 6
8 4
9 4
dtype: int64
In [70]: s.value_counts()
Out[70]:
4 5
2 2
6 2
1 1
dtype: int64
String Methods#
Series is equipped with a set of string processing methods in the str
attribute that make it easy to operate on each element of the array, as in the
code snippet below. Note that pattern-matching in str generally uses regular
expressions by default (and in
some cases always uses them). See more at Vectorized String Methods.
In [71]: s = pd.Series(["A", "B", "C", "Aaba", "Baca", np.nan, "CABA", "dog", "cat"])
In [72]: s.str.lower()
Out[72]:
0 a
1 b
2 c
3 aaba
4 baca
5 NaN
6 caba
7 dog
8 cat
dtype: object
Merge#
Concat#
pandas provides various facilities for easily combining together Series and DataFrame objects with various kinds of set logic for the indexes and relational algebra functionality in the case of join / merge-type operations.
See the Merging section.
Concatenating pandas objects together along an axis with concat():
In [73]: df = pd.DataFrame(np.random.randn(10, 4))
In [74]: df
Out[74]:
0 1 2 3
0 -0.548702 1.467327 -1.015962 -0.483075
1 1.637550 -1.217659 -0.291519 -1.745505
2 -0.263952 0.991460 -0.919069 0.266046
3 -0.709661 1.669052 1.037882 -1.705775
4 -0.919854 -0.042379 1.247642 -0.009920
5 0.290213 0.495767 0.362949 1.548106
6 -1.131345 -0.089329 0.337863 -0.945867
7 -0.932132 1.956030 0.017587 -0.016692
8 -0.575247 0.254161 -1.143704 0.215897
9 1.193555 -0.077118 -0.408530 -0.862495
# break it into pieces
In [75]: pieces = [df[:3], df[3:7], df[7:]]
In [76]: pd.concat(pieces)
Out[76]:
0 1 2 3
0 -0.548702 1.467327 -1.015962 -0.483075
1 1.637550 -1.217659 -0.291519 -1.745505
2 -0.263952 0.991460 -0.919069 0.266046
3 -0.709661 1.669052 1.037882 -1.705775
4 -0.919854 -0.042379 1.247642 -0.009920
5 0.290213 0.495767 0.362949 1.548106
6 -1.131345 -0.089329 0.337863 -0.945867
7 -0.932132 1.956030 0.017587 -0.016692
8 -0.575247 0.254161 -1.143704 0.215897
9 1.193555 -0.077118 -0.408530 -0.862495
Join#
merge() enables SQL style join types along specific columns. See the Database style joining section.
In [77]: left = pd.DataFrame({"key": ["foo", "foo"], "lval": [1, 2]})
In [78]: right = pd.DataFrame({"key": ["foo", "foo"], "rval": [4, 5]})
In [79]: left
Out[79]:
key lval
0 foo 1
1 foo 2
In [80]: right
Out[80]:
key rval
0 foo 4
1 foo 5
In [81]: pd.merge(left, right, on="key")
Out[81]:
key lval rval
0 foo 1 4
1 foo 1 5
2 foo 2 4
3 foo 2 5
Another example that can be given is:
In [82]: left = pd.DataFrame({"key": ["foo", "bar"], "lval": [1, 2]})
In [83]: right = pd.DataFrame({"key": ["foo", "bar"], "rval": [4, 5]})
In [84]: left
Out[84]:
key lval
0 foo 1
1 bar 2
In [85]: right
Out[85]:
key rval
0 foo 4
1 bar 5
In [86]: pd.merge(left, right, on="key")
Out[86]:
key lval rval
0 foo 1 4
1 bar 2 5
Grouping#
By “group by” we are referring to a process involving one or more of the following steps:
Splitting the data into groups based on some criteria
Applying a function to each group independently
Combining the results into a data structure
See the Grouping section.
In [87]: df = pd.DataFrame(
....: {
....: "A": ["foo", "bar", "foo", "bar", "foo", "bar", "foo", "foo"],
....: "B": ["one", "one", "two", "three", "two", "two", "one", "three"],
....: "C": np.random.randn(8),
....: "D": np.random.randn(8),
....: }
....: )
....:
In [88]: df
Out[88]:
A B C D
0 foo one 1.346061 -1.577585
1 bar one 1.511763 0.396823
2 foo two 1.627081 -0.105381
3 bar three -0.990582 -0.532532
4 foo two -0.441652 1.453749
5 bar two 1.211526 1.208843
6 foo one 0.268520 -0.080952
7 foo three 0.024580 -0.264610
Grouping and then applying the sum() function to the resulting
groups:
In [89]: df.groupby("A")[["C", "D"]].sum()
Out[89]:
C D
A
bar 1.732707 1.073134
foo 2.824590 -0.574779
Grouping by multiple columns forms a hierarchical index, and again we can
apply the sum() function:
In [90]: df.groupby(["A", "B"]).sum()
Out[90]:
C D
A B
bar one 1.511763 0.396823
three -0.990582 -0.532532
two 1.211526 1.208843
foo one 1.614581 -1.658537
three 0.024580 -0.264610
two 1.185429 1.348368
Reshaping#
See the sections on Hierarchical Indexing and Reshaping.
Stack#
In [91]: tuples = list(
....: zip(
....: ["bar", "bar", "baz", "baz", "foo", "foo", "qux", "qux"],
....: ["one", "two", "one", "two", "one", "two", "one", "two"],
....: )
....: )
....:
In [92]: index = pd.MultiIndex.from_tuples(tuples, names=["first", "second"])
In [93]: df = pd.DataFrame(np.random.randn(8, 2), index=index, columns=["A", "B"])
In [94]: df2 = df[:4]
In [95]: df2
Out[95]:
A B
first second
bar one -0.727965 -0.589346
two 0.339969 -0.693205
baz one -0.339355 0.593616
two 0.884345 1.591431
The stack() method “compresses” a level in the DataFrame’s
columns:
In [96]: stacked = df2.stack()
In [97]: stacked
Out[97]:
first second
bar one A -0.727965
B -0.589346
two A 0.339969
B -0.693205
baz one A -0.339355
B 0.593616
two A 0.884345
B 1.591431
dtype: float64
With a “stacked” DataFrame or Series (having a MultiIndex as the
index), the inverse operation of stack() is
unstack(), which by default unstacks the last level:
In [98]: stacked.unstack()
Out[98]:
A B
first second
bar one -0.727965 -0.589346
two 0.339969 -0.693205
baz one -0.339355 0.593616
two 0.884345 1.591431
In [99]: stacked.unstack(1)
Out[99]:
second one two
first
bar A -0.727965 0.339969
B -0.589346 -0.693205
baz A -0.339355 0.884345
B 0.593616 1.591431
In [100]: stacked.unstack(0)
Out[100]:
first bar baz
second
one A -0.727965 -0.339355
B -0.589346 0.593616
two A 0.339969 0.884345
B -0.693205 1.591431
Pivot tables#
See the section on Pivot Tables.
In [101]: df = pd.DataFrame(
.....: {
.....: "A": ["one", "one", "two", "three"] * 3,
.....: "B": ["A", "B", "C"] * 4,
.....: "C": ["foo", "foo", "foo", "bar", "bar", "bar"] * 2,
.....: "D": np.random.randn(12),
.....: "E": np.random.randn(12),
.....: }
.....: )
.....:
In [102]: df
Out[102]:
A B C D E
0 one A foo -1.202872 0.047609
1 one B foo -1.814470 -0.136473
2 two C foo 1.018601 -0.561757
3 three A bar -0.595447 -1.623033
4 one B bar 1.395433 0.029399
5 one C bar -0.392670 -0.542108
6 two A foo 0.007207 0.282696
7 three B foo 1.928123 -0.087302
8 one C foo -0.055224 -1.575170
9 one A bar 2.395985 1.771208
10 two B bar 1.552825 0.816482
11 three C bar 0.166599 1.100230
pivot_table() pivots a DataFrame specifying the values, index and columns
In [103]: pd.pivot_table(df, values="D", index=["A", "B"], columns=["C"])
Out[103]:
C bar foo
A B
one A 2.395985 -1.202872
B 1.395433 -1.814470
C -0.392670 -0.055224
three A -0.595447 NaN
B NaN 1.928123
C 0.166599 NaN
two A NaN 0.007207
B 1.552825 NaN
C NaN 1.018601
Time series#
pandas has simple, powerful, and efficient functionality for performing resampling operations during frequency conversion (e.g., converting secondly data into 5-minutely data). This is extremely common in, but not limited to, financial applications. See the Time Series section.
In [104]: rng = pd.date_range("1/1/2012", periods=100, freq="S")
In [105]: ts = pd.Series(np.random.randint(0, 500, len(rng)), index=rng)
In [106]: ts.resample("5Min").sum()
Out[106]:
2012-01-01 24182
Freq: 5T, dtype: int64
Series.tz_localize() localizes a time series to a time zone:
In [107]: rng = pd.date_range("3/6/2012 00:00", periods=5, freq="D")
In [108]: ts = pd.Series(np.random.randn(len(rng)), rng)
In [109]: ts
Out[109]:
2012-03-06 1.857704
2012-03-07 -1.193545
2012-03-08 0.677510
2012-03-09 -0.153931
2012-03-10 0.520091
Freq: D, dtype: float64
In [110]: ts_utc = ts.tz_localize("UTC")
In [111]: ts_utc
Out[111]:
2012-03-06 00:00:00+00:00 1.857704
2012-03-07 00:00:00+00:00 -1.193545
2012-03-08 00:00:00+00:00 0.677510
2012-03-09 00:00:00+00:00 -0.153931
2012-03-10 00:00:00+00:00 0.520091
Freq: D, dtype: float64
Series.tz_convert() converts a timezones aware time series to another time zone:
In [112]: ts_utc.tz_convert("US/Eastern")
Out[112]:
2012-03-05 19:00:00-05:00 1.857704
2012-03-06 19:00:00-05:00 -1.193545
2012-03-07 19:00:00-05:00 0.677510
2012-03-08 19:00:00-05:00 -0.153931
2012-03-09 19:00:00-05:00 0.520091
Freq: D, dtype: float64
Converting between time span representations:
In [113]: rng = pd.date_range("1/1/2012", periods=5, freq="M")
In [114]: ts = pd.Series(np.random.randn(len(rng)), index=rng)
In [115]: ts
Out[115]:
2012-01-31 -1.475051
2012-02-29 0.722570
2012-03-31 -0.322646
2012-04-30 -1.601631
2012-05-31 0.778033
Freq: M, dtype: float64
In [116]: ps = ts.to_period()
In [117]: ps
Out[117]:
2012-01 -1.475051
2012-02 0.722570
2012-03 -0.322646
2012-04 -1.601631
2012-05 0.778033
Freq: M, dtype: float64
In [118]: ps.to_timestamp()
Out[118]:
2012-01-01 -1.475051
2012-02-01 0.722570
2012-03-01 -0.322646
2012-04-01 -1.601631
2012-05-01 0.778033
Freq: MS, dtype: float64
Converting between period and timestamp enables some convenient arithmetic functions to be used. In the following example, we convert a quarterly frequency with year ending in November to 9am of the end of the month following the quarter end:
In [119]: prng = pd.period_range("1990Q1", "2000Q4", freq="Q-NOV")
In [120]: ts = pd.Series(np.random.randn(len(prng)), prng)
In [121]: ts.index = (prng.asfreq("M", "e") + 1).asfreq("H", "s") + 9
In [122]: ts.head()
Out[122]:
1990-03-01 09:00 -0.289342
1990-06-01 09:00 0.233141
1990-09-01 09:00 -0.223540
1990-12-01 09:00 0.542054
1991-03-01 09:00 -0.688585
Freq: H, dtype: float64
Categoricals#
pandas can include categorical data in a DataFrame. For full docs, see the
categorical introduction and the API documentation.
In [123]: df = pd.DataFrame(
.....: {"id": [1, 2, 3, 4, 5, 6], "raw_grade": ["a", "b", "b", "a", "a", "e"]}
.....: )
.....:
Converting the raw grades to a categorical data type:
In [124]: df["grade"] = df["raw_grade"].astype("category")
In [125]: df["grade"]
Out[125]:
0 a
1 b
2 b
3 a
4 a
5 e
Name: grade, dtype: category
Categories (3, object): ['a', 'b', 'e']
Rename the categories to more meaningful names:
In [126]: new_categories = ["very good", "good", "very bad"]
In [127]: df["grade"] = df["grade"].cat.rename_categories(new_categories)
Reorder the categories and simultaneously add the missing categories (methods under Series.cat() return a new Series by default):
In [128]: df["grade"] = df["grade"].cat.set_categories(
.....: ["very bad", "bad", "medium", "good", "very good"]
.....: )
.....:
In [129]: df["grade"]
Out[129]:
0 very good
1 good
2 good
3 very good
4 very good
5 very bad
Name: grade, dtype: category
Categories (5, object): ['very bad', 'bad', 'medium', 'good', 'very good']
Sorting is per order in the categories, not lexical order:
In [130]: df.sort_values(by="grade")
Out[130]:
id raw_grade grade
5 6 e very bad
1 2 b good
2 3 b good
0 1 a very good
3 4 a very good
4 5 a very good
Grouping by a categorical column also shows empty categories:
In [131]: df.groupby("grade").size()
Out[131]:
grade
very bad 1
bad 0
medium 0
good 2
very good 3
dtype: int64
Plotting#
See the Plotting docs.
We use the standard convention for referencing the matplotlib API:
In [132]: import matplotlib.pyplot as plt
In [133]: plt.close("all")
The plt.close method is used to close a figure window:
In [134]: ts = pd.Series(np.random.randn(1000), index=pd.date_range("1/1/2000", periods=1000))
In [135]: ts = ts.cumsum()
In [136]: ts.plot();
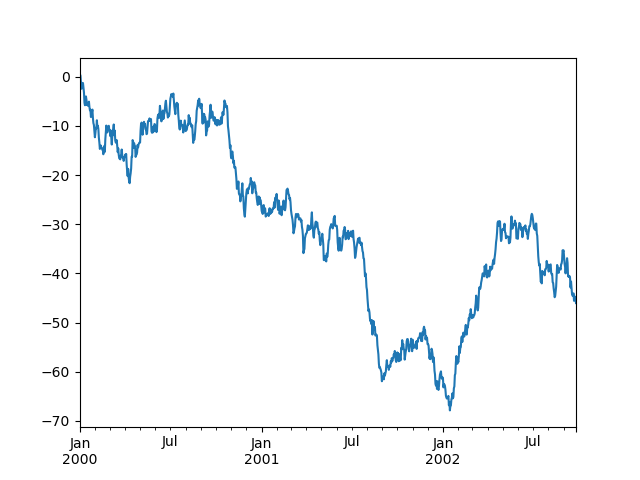
If running under Jupyter Notebook, the plot will appear on plot(). Otherwise use
matplotlib.pyplot.show to show it or
matplotlib.pyplot.savefig to write it to a file.
In [137]: plt.show();
On a DataFrame, the plot() method is a convenience to plot all
of the columns with labels:
In [138]: df = pd.DataFrame(
.....: np.random.randn(1000, 4), index=ts.index, columns=["A", "B", "C", "D"]
.....: )
.....:
In [139]: df = df.cumsum()
In [140]: plt.figure();
In [141]: df.plot();
In [142]: plt.legend(loc='best');
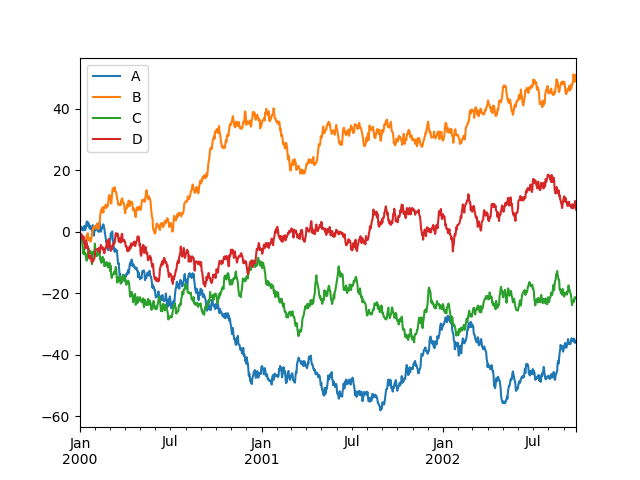
Importing and exporting data#
CSV#
Writing to a csv file: using DataFrame.to_csv()
In [143]: df.to_csv("foo.csv")
Reading from a csv file: using read_csv()
In [144]: pd.read_csv("foo.csv")
Out[144]:
Unnamed: 0 A B C D
0 2000-01-01 0.350262 0.843315 1.798556 0.782234
1 2000-01-02 -0.586873 0.034907 1.923792 -0.562651
2 2000-01-03 -1.245477 -0.963406 2.269575 -1.612566
3 2000-01-04 -0.252830 -0.498066 3.176886 -1.275581
4 2000-01-05 -1.044057 0.118042 2.768571 0.386039
.. ... ... ... ... ...
995 2002-09-22 -48.017654 31.474551 69.146374 -47.541670
996 2002-09-23 -47.207912 32.627390 68.505254 -48.828331
997 2002-09-24 -48.907133 31.990402 67.310924 -49.391051
998 2002-09-25 -50.146062 33.716770 67.717434 -49.037577
999 2002-09-26 -49.724318 33.479952 68.108014 -48.822030
[1000 rows x 5 columns]
HDF5#
Reading and writing to HDFStores.
Writing to a HDF5 Store using DataFrame.to_hdf():
In [145]: df.to_hdf("foo.h5", "df")
Reading from a HDF5 Store using read_hdf():
In [146]: pd.read_hdf("foo.h5", "df")
Out[146]:
A B C D
2000-01-01 0.350262 0.843315 1.798556 0.782234
2000-01-02 -0.586873 0.034907 1.923792 -0.562651
2000-01-03 -1.245477 -0.963406 2.269575 -1.612566
2000-01-04 -0.252830 -0.498066 3.176886 -1.275581
2000-01-05 -1.044057 0.118042 2.768571 0.386039
... ... ... ... ...
2002-09-22 -48.017654 31.474551 69.146374 -47.541670
2002-09-23 -47.207912 32.627390 68.505254 -48.828331
2002-09-24 -48.907133 31.990402 67.310924 -49.391051
2002-09-25 -50.146062 33.716770 67.717434 -49.037577
2002-09-26 -49.724318 33.479952 68.108014 -48.822030
[1000 rows x 4 columns]
Excel#
Reading and writing to Excel.
Writing to an excel file using DataFrame.to_excel():
In [147]: df.to_excel("foo.xlsx", sheet_name="Sheet1")
Reading from an excel file using read_excel():
In [148]: pd.read_excel("foo.xlsx", "Sheet1", index_col=None, na_values=["NA"])
Out[148]:
Unnamed: 0 A B C D
0 2000-01-01 0.350262 0.843315 1.798556 0.782234
1 2000-01-02 -0.586873 0.034907 1.923792 -0.562651
2 2000-01-03 -1.245477 -0.963406 2.269575 -1.612566
3 2000-01-04 -0.252830 -0.498066 3.176886 -1.275581
4 2000-01-05 -1.044057 0.118042 2.768571 0.386039
.. ... ... ... ... ...
995 2002-09-22 -48.017654 31.474551 69.146374 -47.541670
996 2002-09-23 -47.207912 32.627390 68.505254 -48.828331
997 2002-09-24 -48.907133 31.990402 67.310924 -49.391051
998 2002-09-25 -50.146062 33.716770 67.717434 -49.037577
999 2002-09-26 -49.724318 33.479952 68.108014 -48.822030
[1000 rows x 5 columns]
Gotchas#
If you are attempting to perform a boolean operation on a Series or DataFrame
you might see an exception like:
In [149]: if pd.Series([False, True, False]):
.....: print("I was true")
.....:
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[149], line 1
----> 1 if pd.Series([False, True, False]):
2 print("I was true")
File ~/work/pandas/pandas/pandas/core/generic.py:1527, in NDFrame.__nonzero__(self)
1525 @final
1526 def __nonzero__(self) -> NoReturn:
-> 1527 raise ValueError(
1528 f"The truth value of a {type(self).__name__} is ambiguous. "
1529 "Use a.empty, a.bool(), a.item(), a.any() or a.all()."
1530 )
ValueError: The truth value of a Series is ambiguous. Use a.empty, a.bool(), a.item(), a.any() or a.all().
See Comparisons and Gotchas for an explanation and what to do.